Dynamics of aptamers
Leveraging NMR data to characterize the conformational landscape of aptamers
ARNA, INSERM U1212, CNRS UMR 5320, Université de Bordeaux
UFR des Sciences Pharmaceutiques, Université de Bordeaux
May 17, 2026
On the limits of mFold
Binding loops stemming from wishful thinking
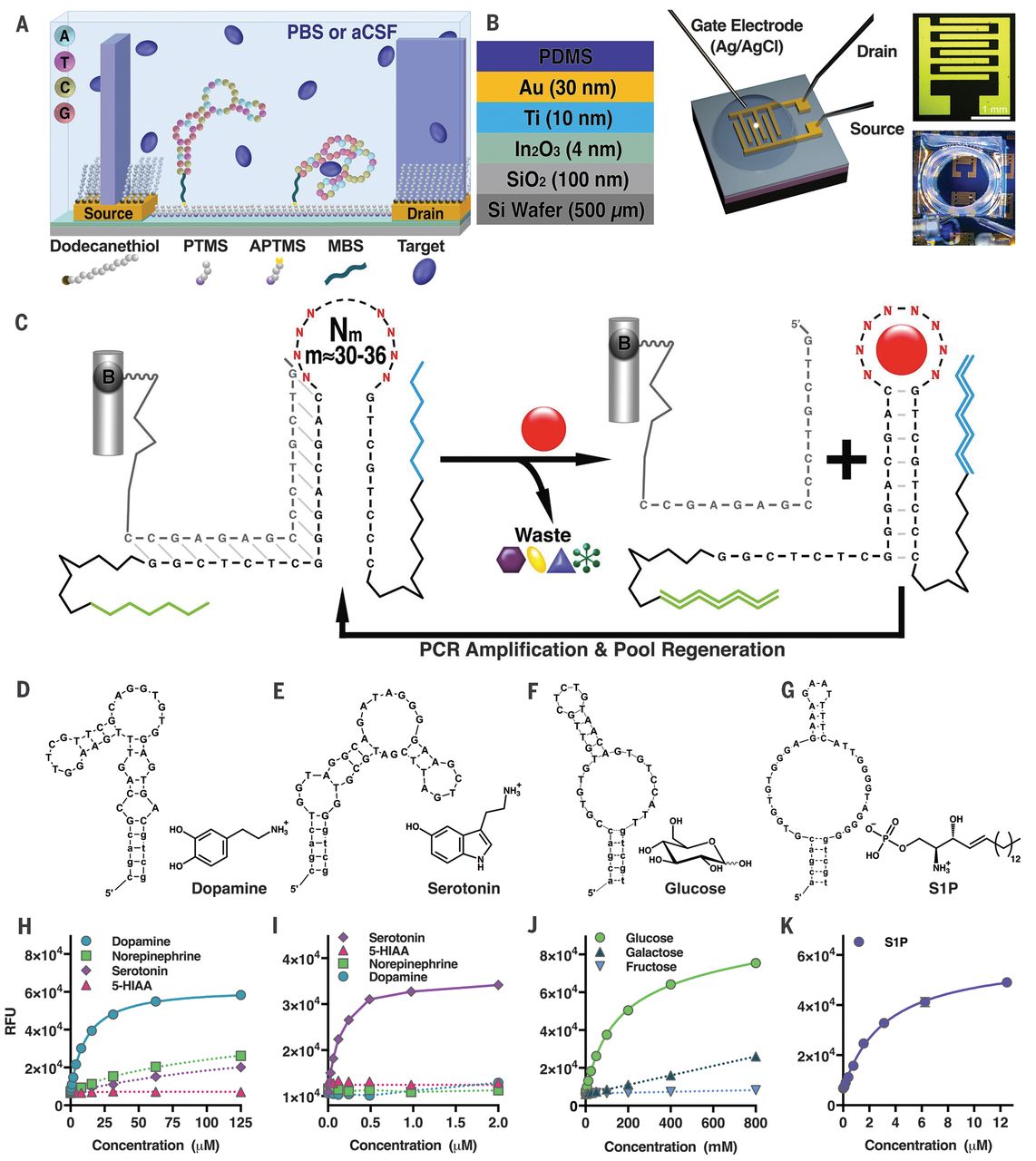
Nakatsuka N., et al., Science, 2018, 362, 319-324
Binding loops stemming from wishful thinking
mFold produces this structure in 1 M NaCl solutions, at 37 °C
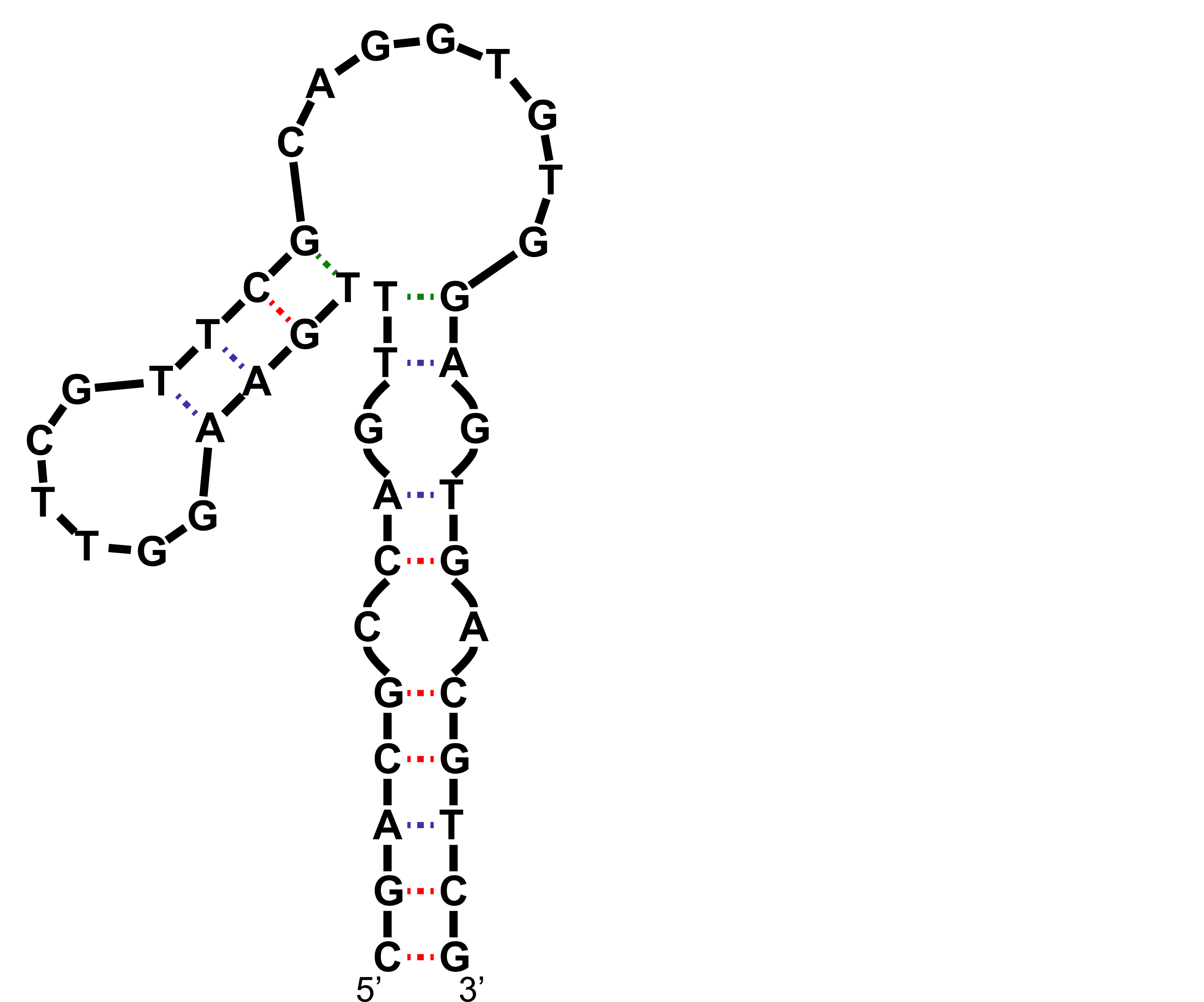
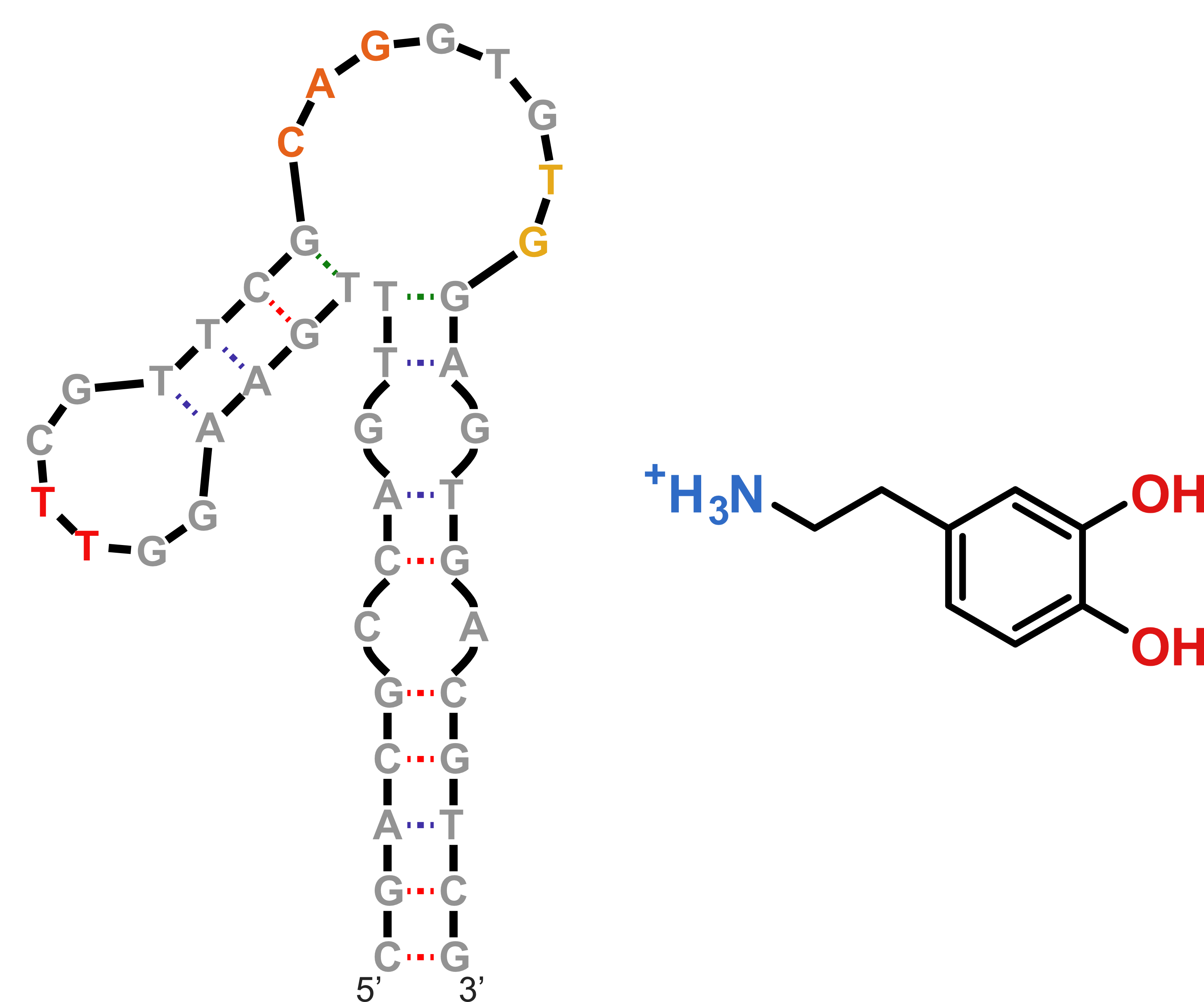
NMR to the rescue
NMR provides a (non-canonical !) starting model
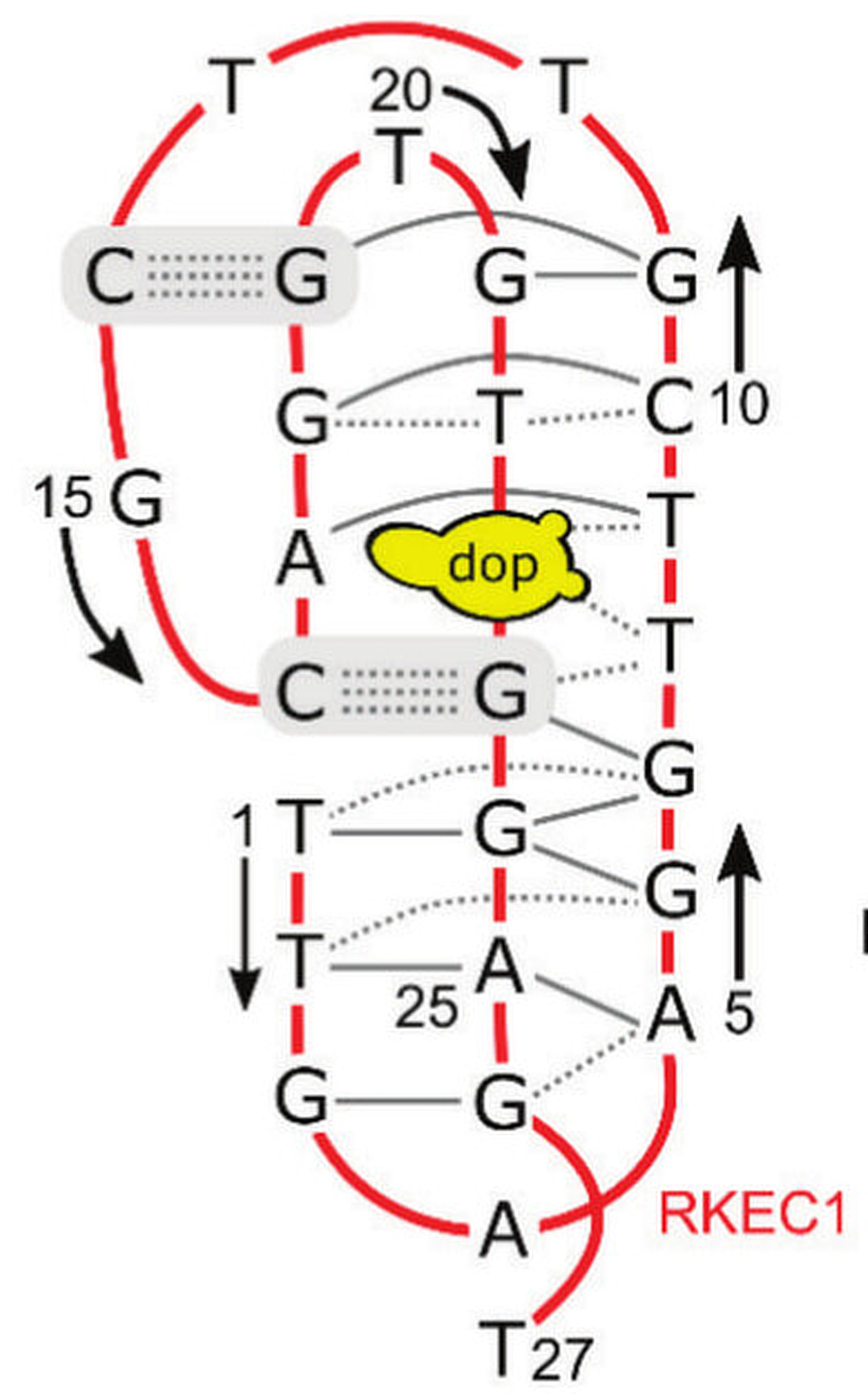
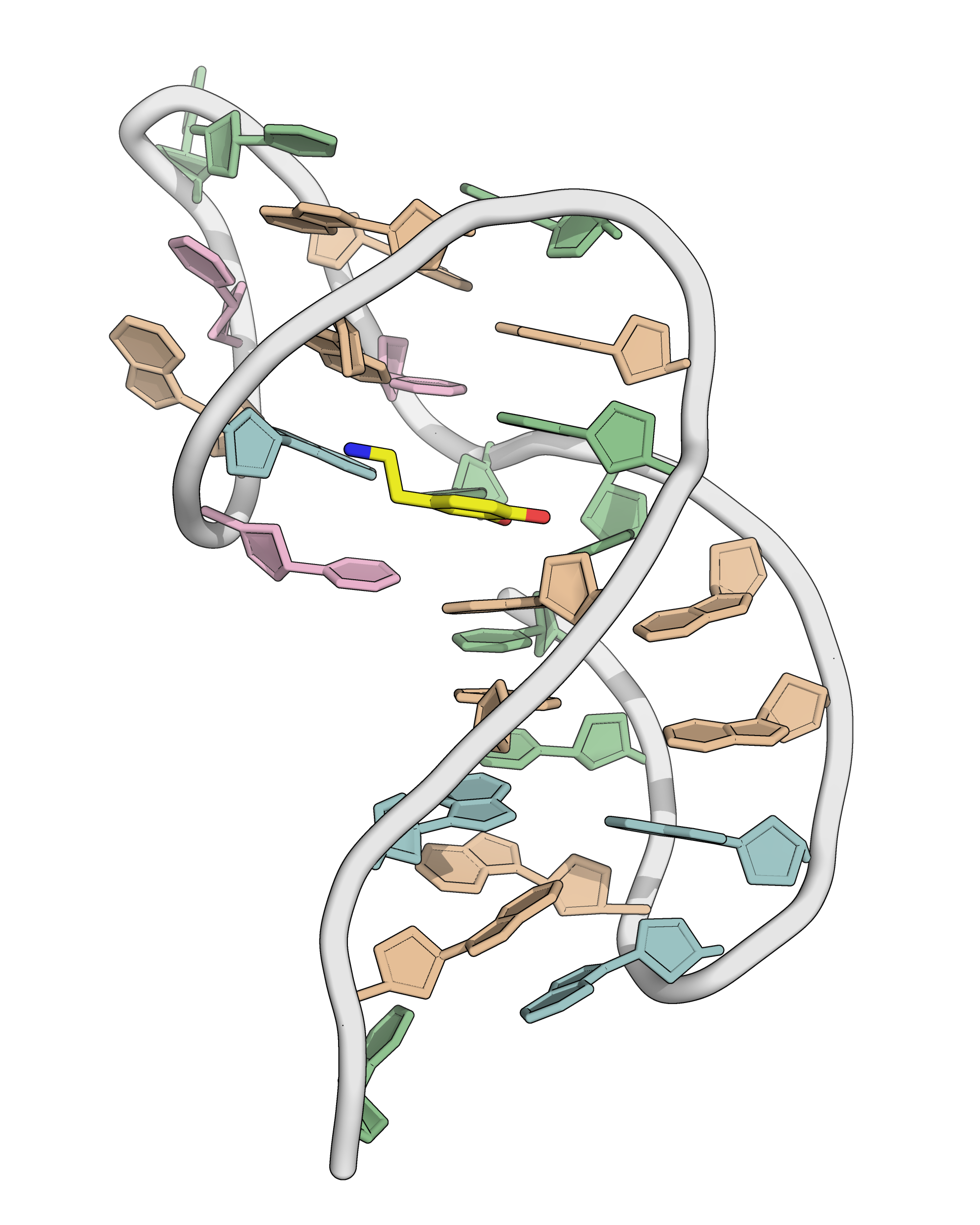
Kretsch R.C., et al., Proteins, 2025
NMR provides an ensemble of models
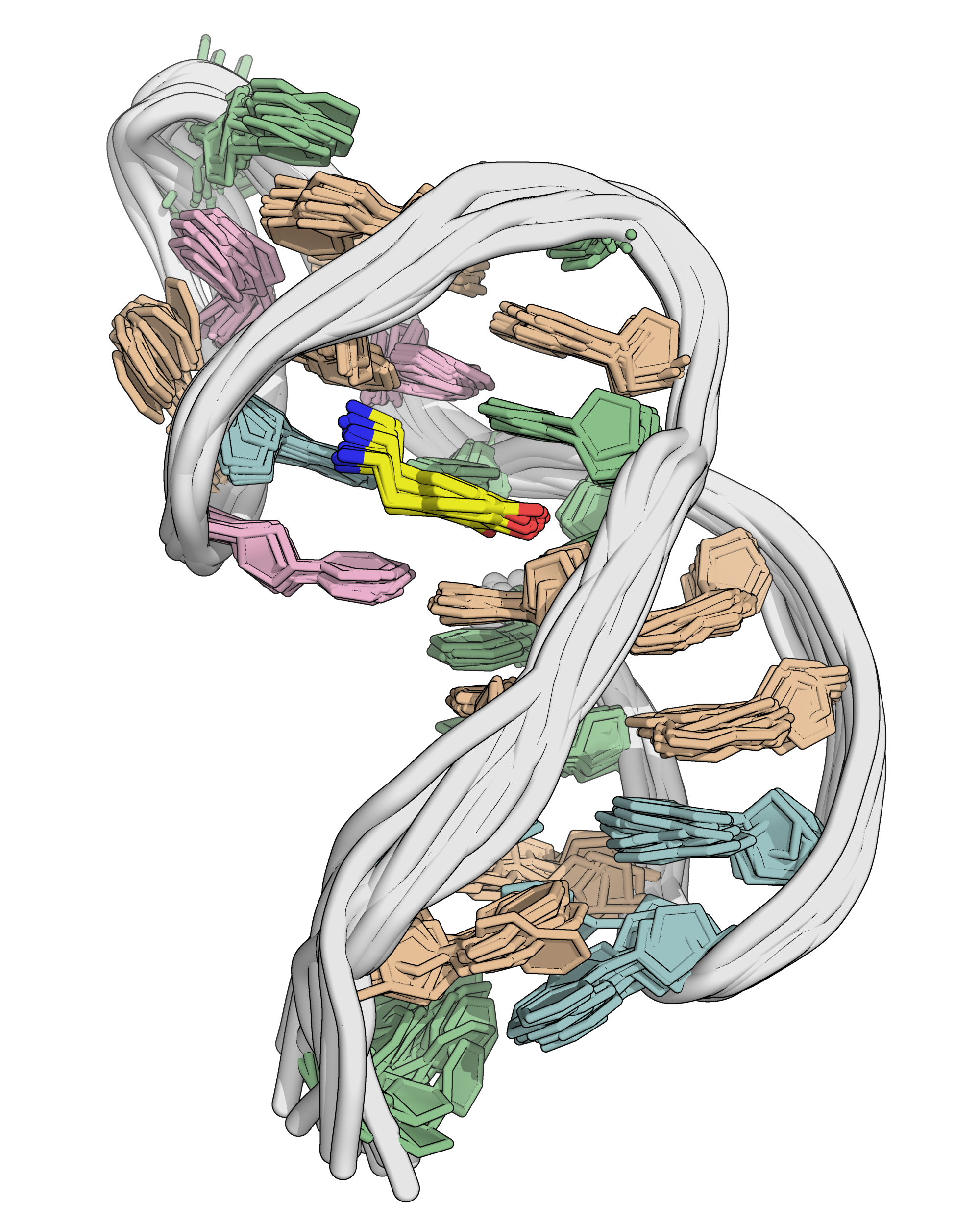
NMR provides many constraints
- DNA (806)
- intra-residue
- inter-residue
- Dopamine (67)
- intramolecular
- intermolecular
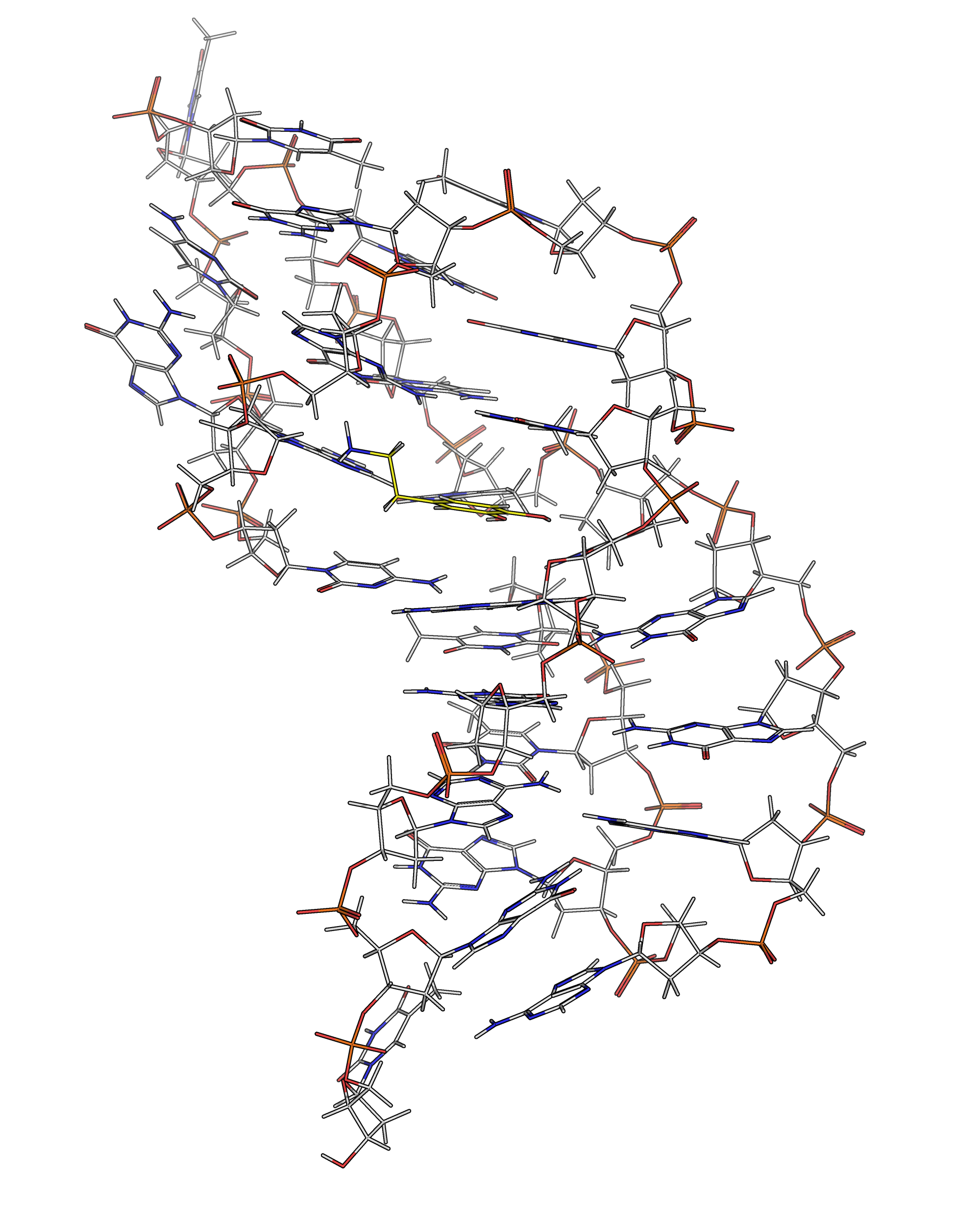
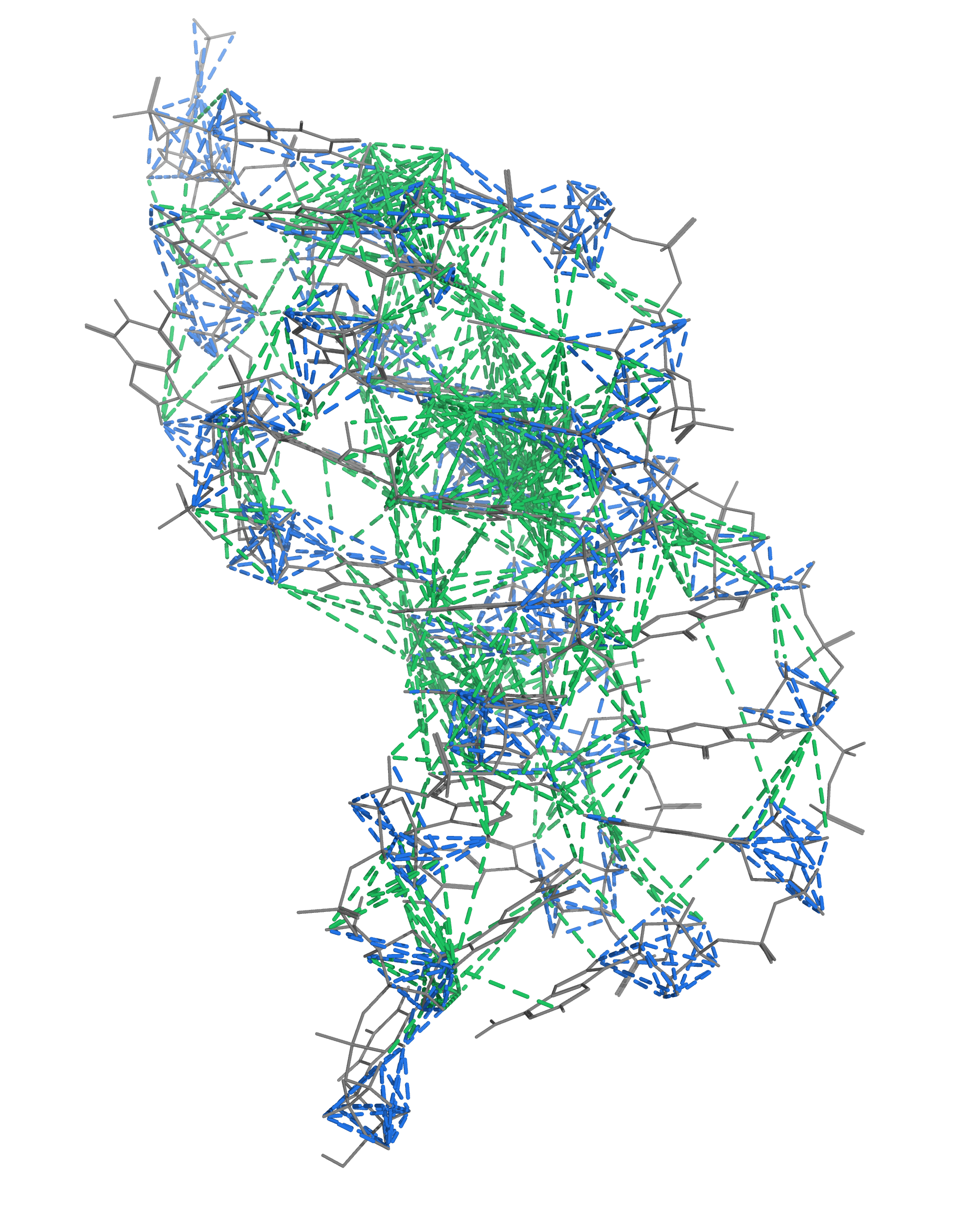
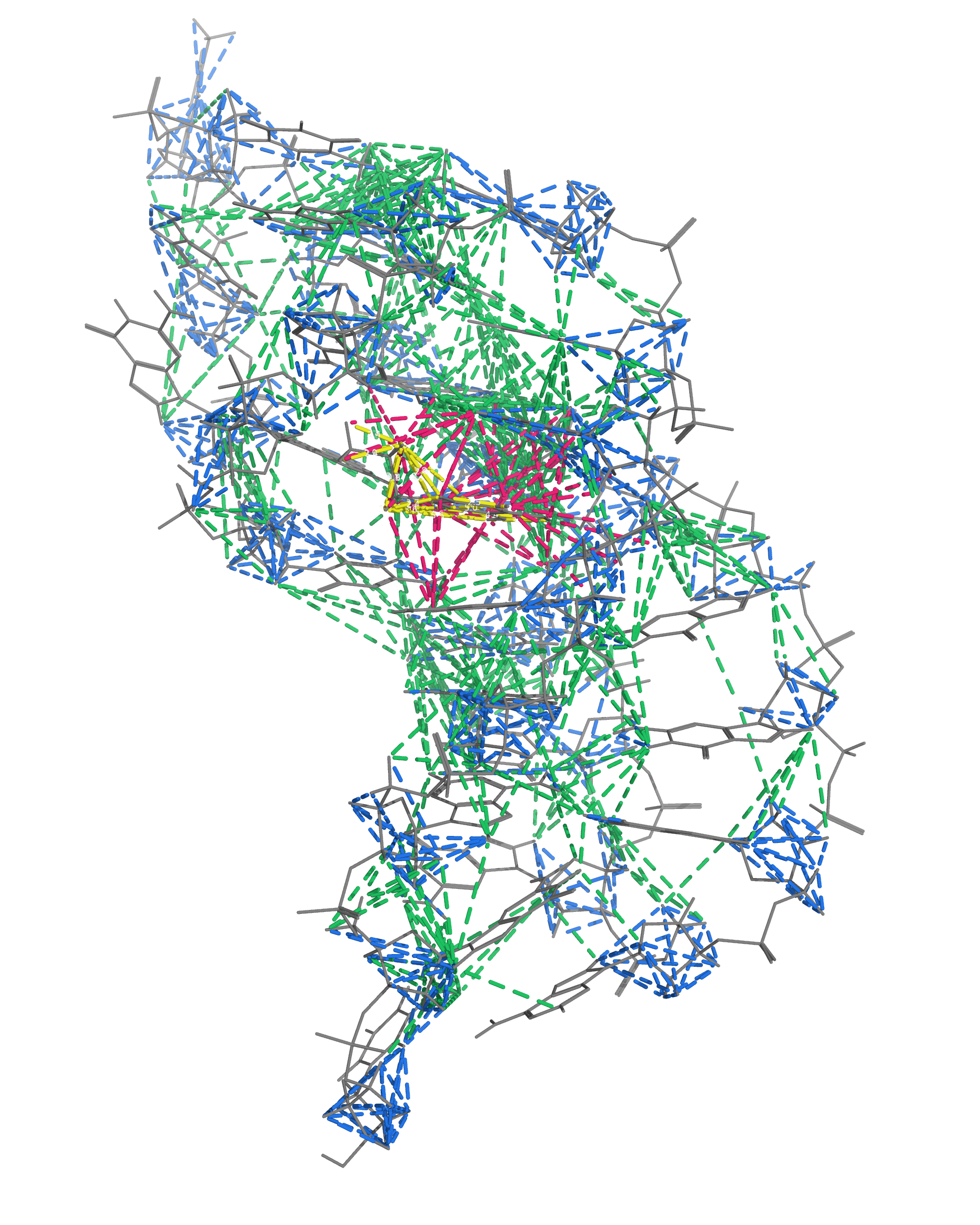
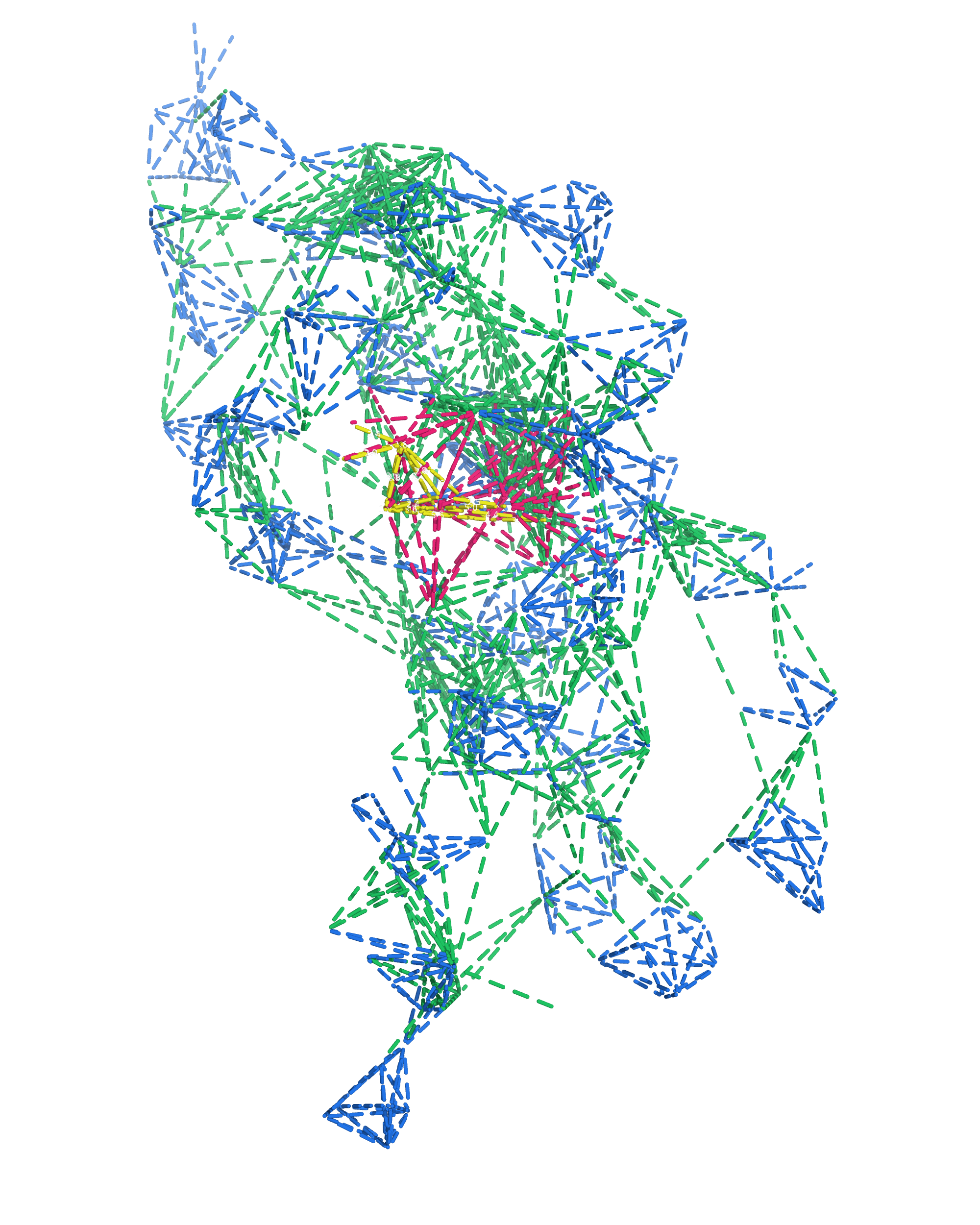
A four step workflow is necessary
- System parameterization for MD
- Force fields, solvation, ions
- System preparation
- Initial relaxation, density correction of mobile molecules first
- Production
- Short runs for minimization of initial models
- Long runs for sampling of conformational space
- Minimization
- Production of an ensemble for PDB and binding site analysis
A four step workflow is necessary
- System parameterization for MD
- Force fields, solvation, ions
- System preparation
- Initial relaxation, density correction of mobile molecules first
- System restraining
- Distance & angles conversion to Amber format
- Force constant tuning for optimal sampling of conformational space
- Production
- Short runs for minimization and longer runs for dynamics sampling
- Minimization
- Production of an ensemble for PDB and binding site analysis
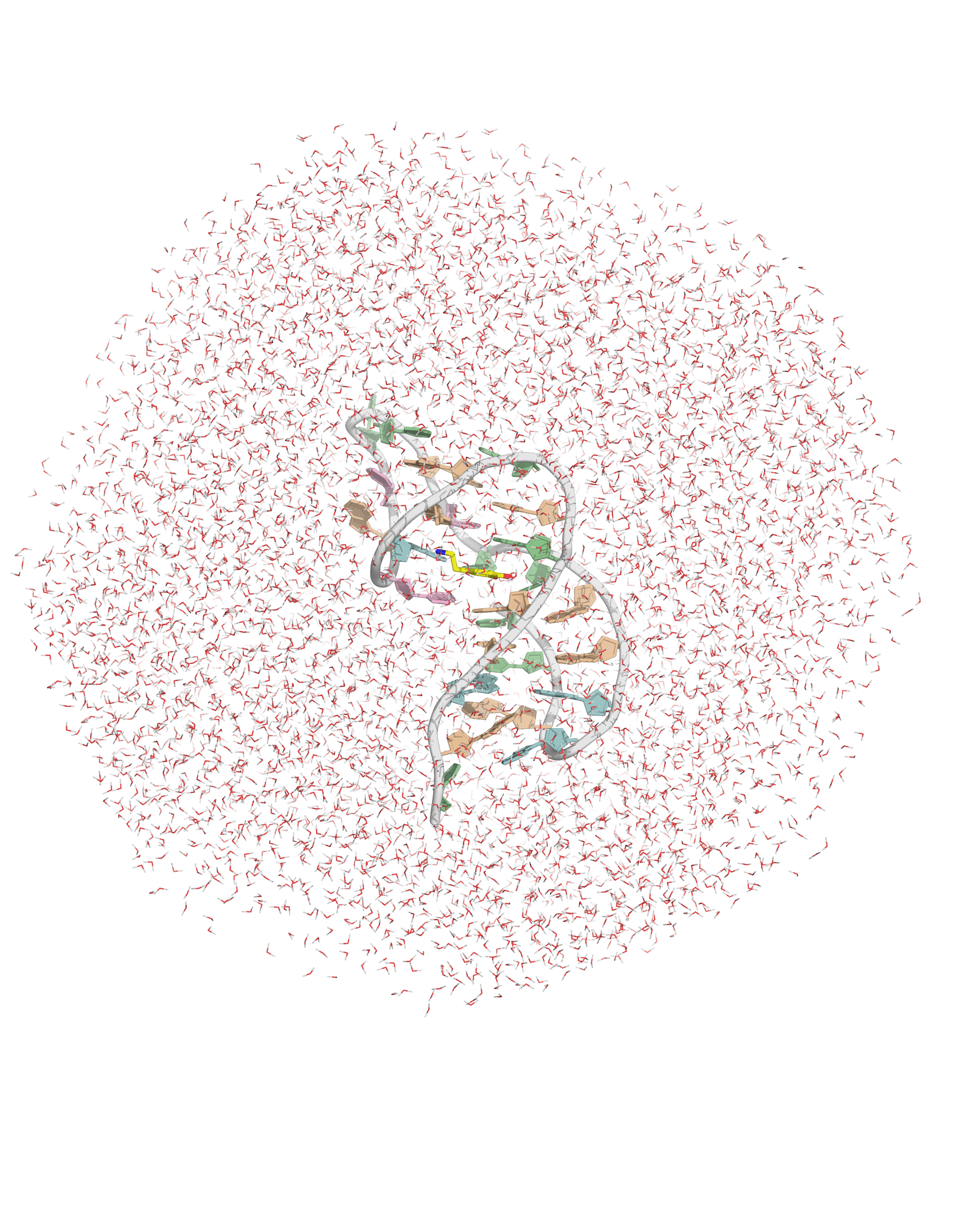
Solvation requires building a box with a model
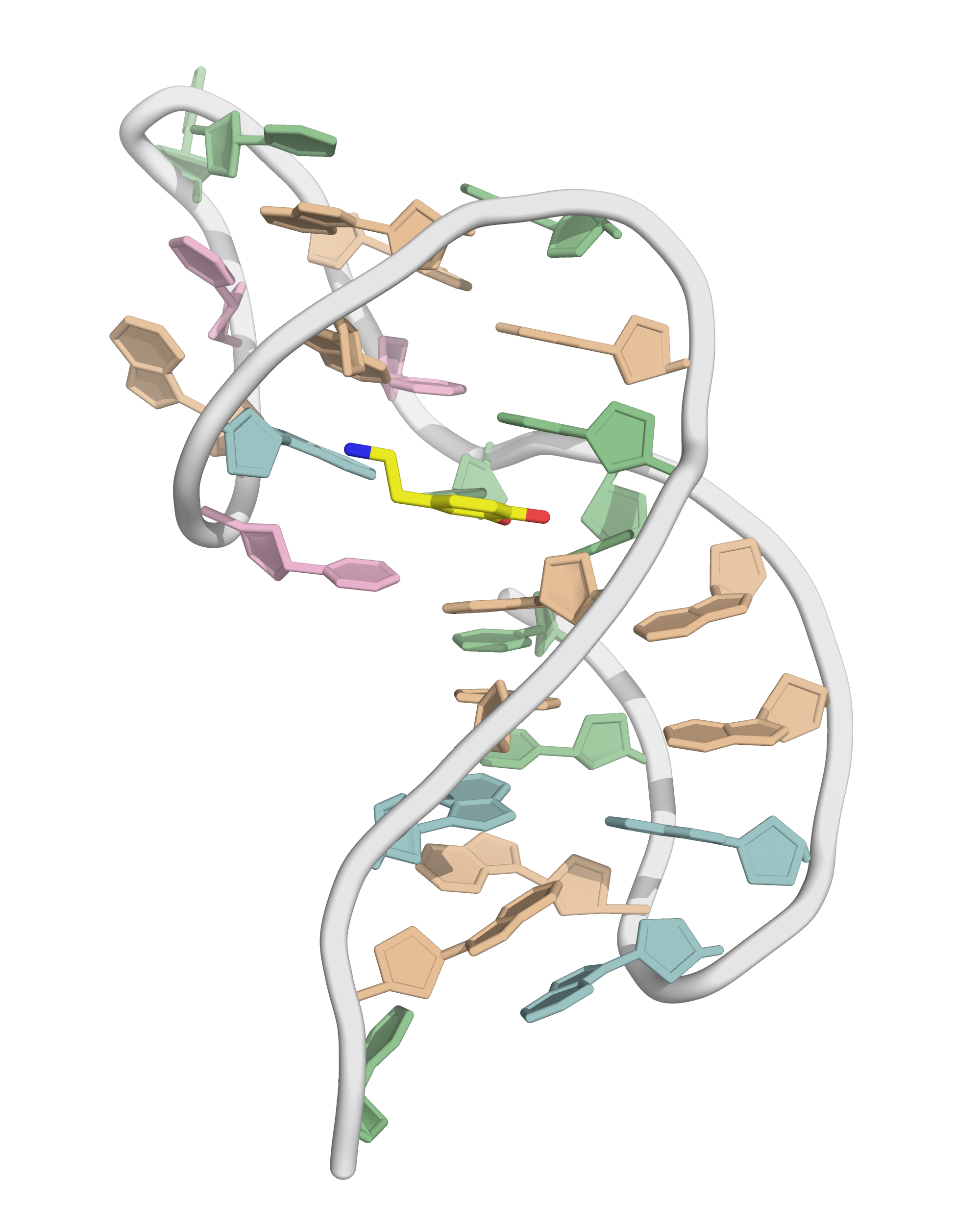
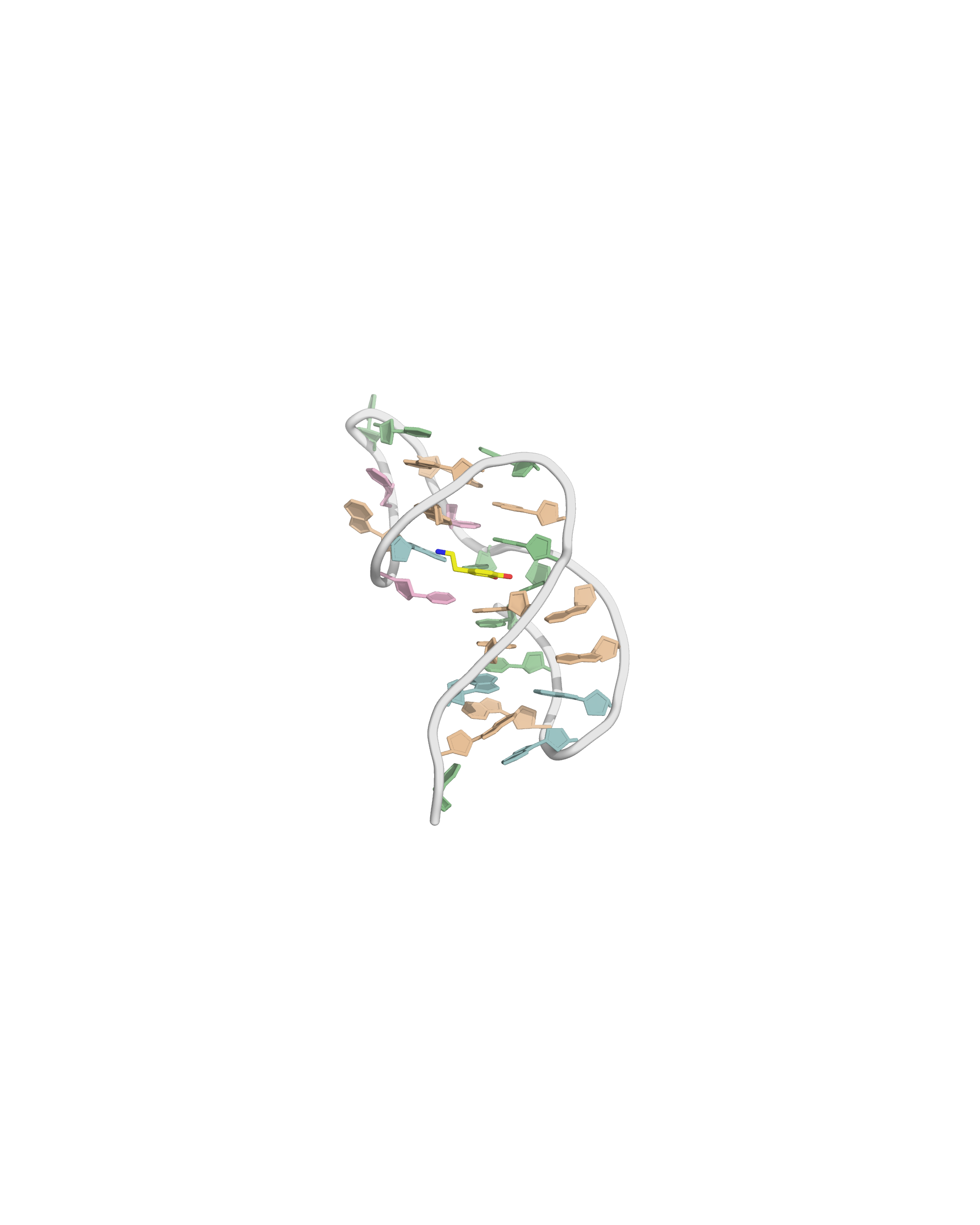
Solvation requires building a box with a model
- OPC water model
- Octahedral box with, 14 Å to surface
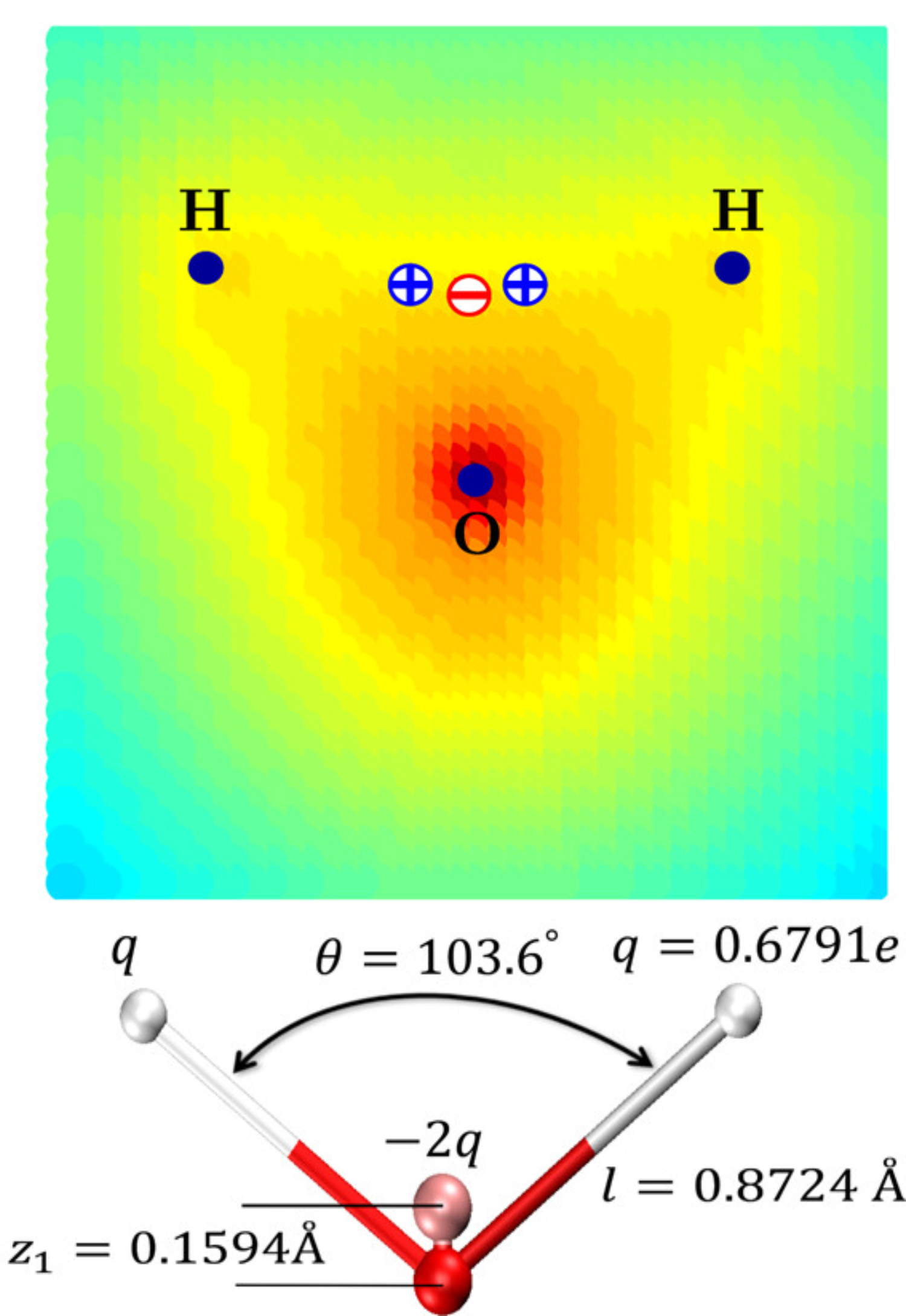

Charge distribution of H2O (gas phase, QM) and 4-point model better reflecting electrostatic potential
Izadi et al., J. Phys. Chem. Lett. 2014, 5, 21, 3863–3871
Na+ ions are added to neutralize the system
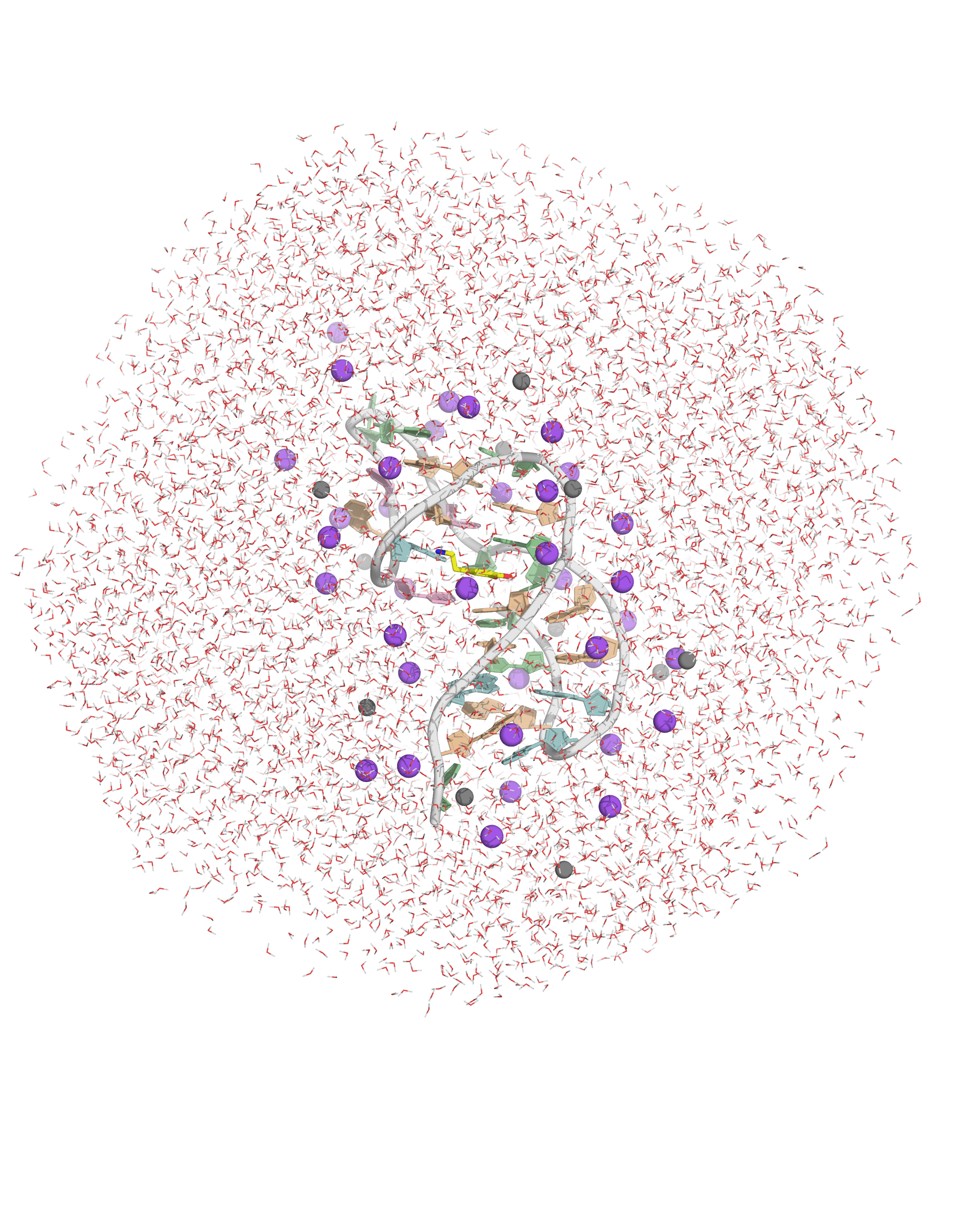
Ions placed near electrostatic potential field minima
Na+ and Cl- are added to mimic experimental conditions
\(N_{\pm}=\color{#376bd3}{\nu_w}\color{#1a9c1a}{c_0}\color{#000000}e^{\mp \operatorname{arcsinh}\!\left(\frac{\color{#b92424}{Q}}{2\color{#376bd3}{\nu_w}\color{#1a9c1a}{c_0}}\right)}\)
- \(\color{#376bd3}{\nu_w}\): water volume of the box (~ 3x105 Å3)
- \(\color{#1a9c1a}{c_0}\): salt concentration (140 mM)
- \(\color{#b92424}{Q}\): charge (-25)

35 Na+ + 10 Cl-
Schmit JD, Kariyawasam NL, Needham V, et al. J. Chem. Theory Comput. 2018;14:1823–1827
Machado MR, Pantano S. J. Chem. Theory Comput. 2020;16:1367–1372
One or several force fields must be used for biomolecules
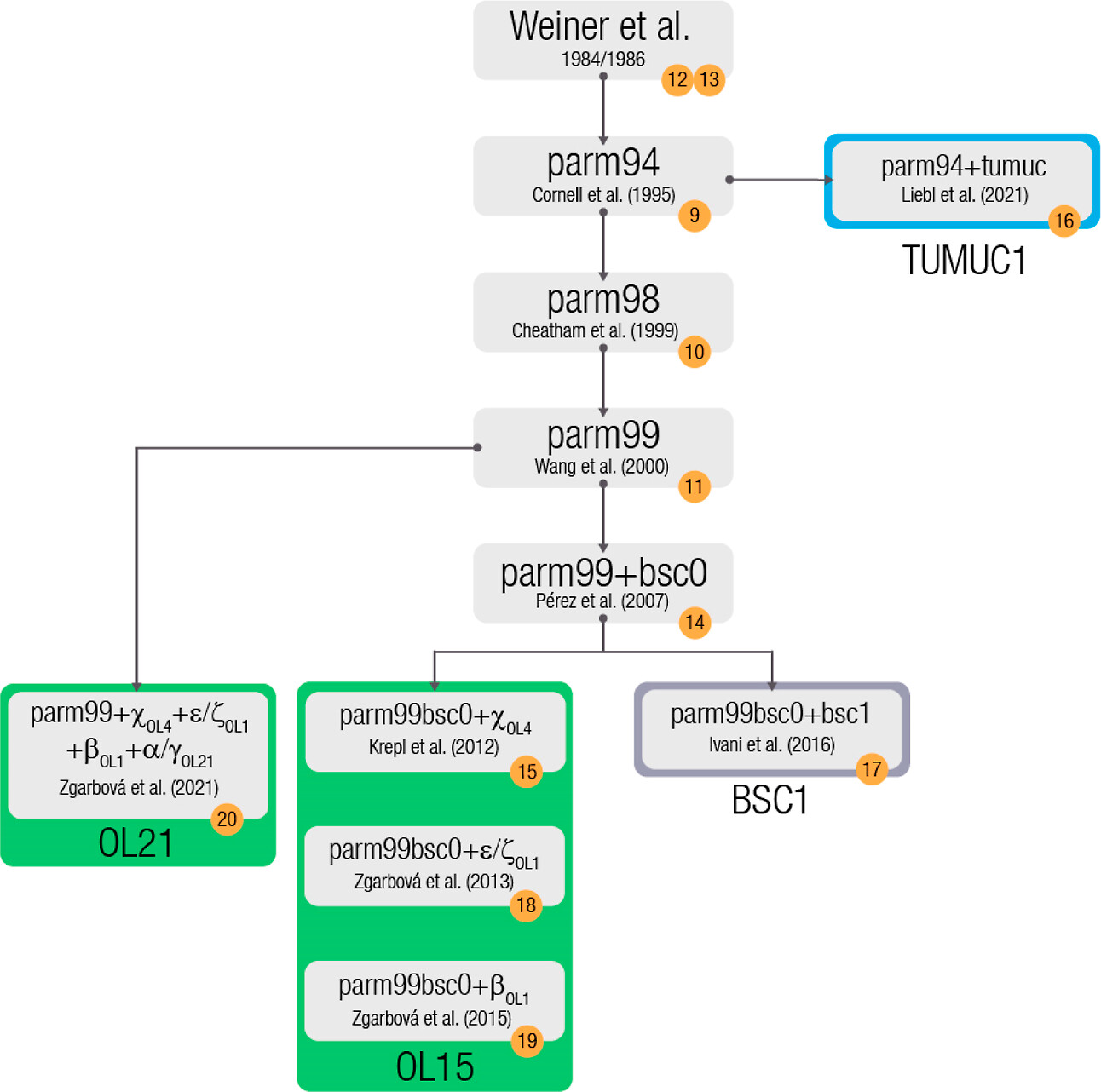
Love O, et al., J. Chem. Theory Comput. 2023, 19, 13, 4299–4307
Where necessary, parameters for ligands must be generated

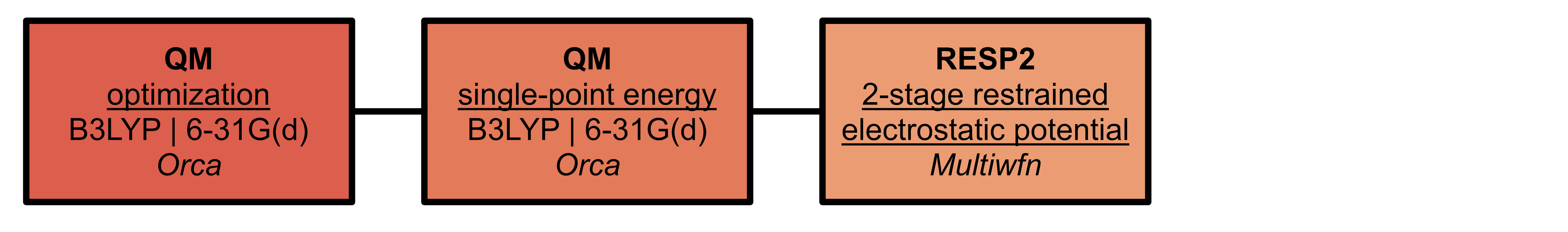
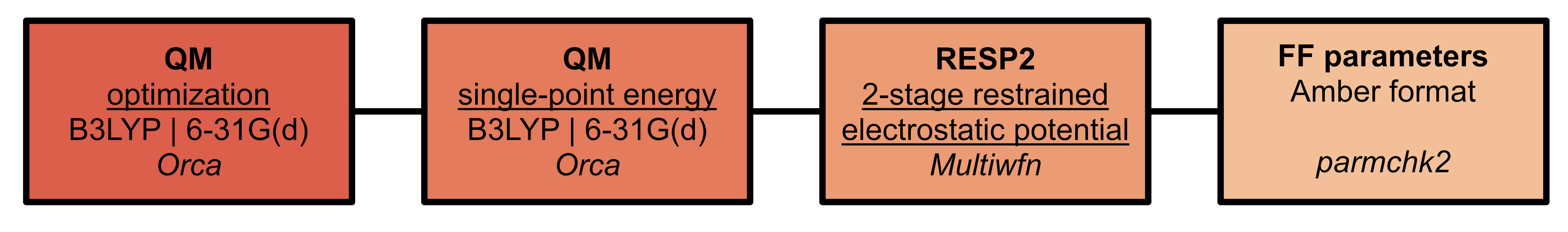
High-performance computing considerations
- Microsecond-scale simulations require GPUs
- Risk of overflow with GPUs for large forces (close contacts)
- CPUs preferred for preparation/minimization steps (double precision)
- GPU and CPU calculations performed with AMBER
- Particle Mesh Ewald Molecular Dynamics (PMEMD) engine, CUDA version for GPUs
- GPU: NVIDIA H100 PCIe Tensor core (~ 40 k€ 💸)
- DOREMI CALI v3 cluster of the MCIA high-performance computing mesocenter (U. Bordeaux, France)
- Simulations ran
- At constant temperature (298 K) and pressure (1 atm)
- With (rMD) and without (uMD) NMR restraints
- 10 ns followed by minimization for PDB deposition
- 1 µs production runs for analysis of the dynamics
Minimized ensemble production by short, restrained MD simulations
10-ns trajectories are stable RMSD-wise
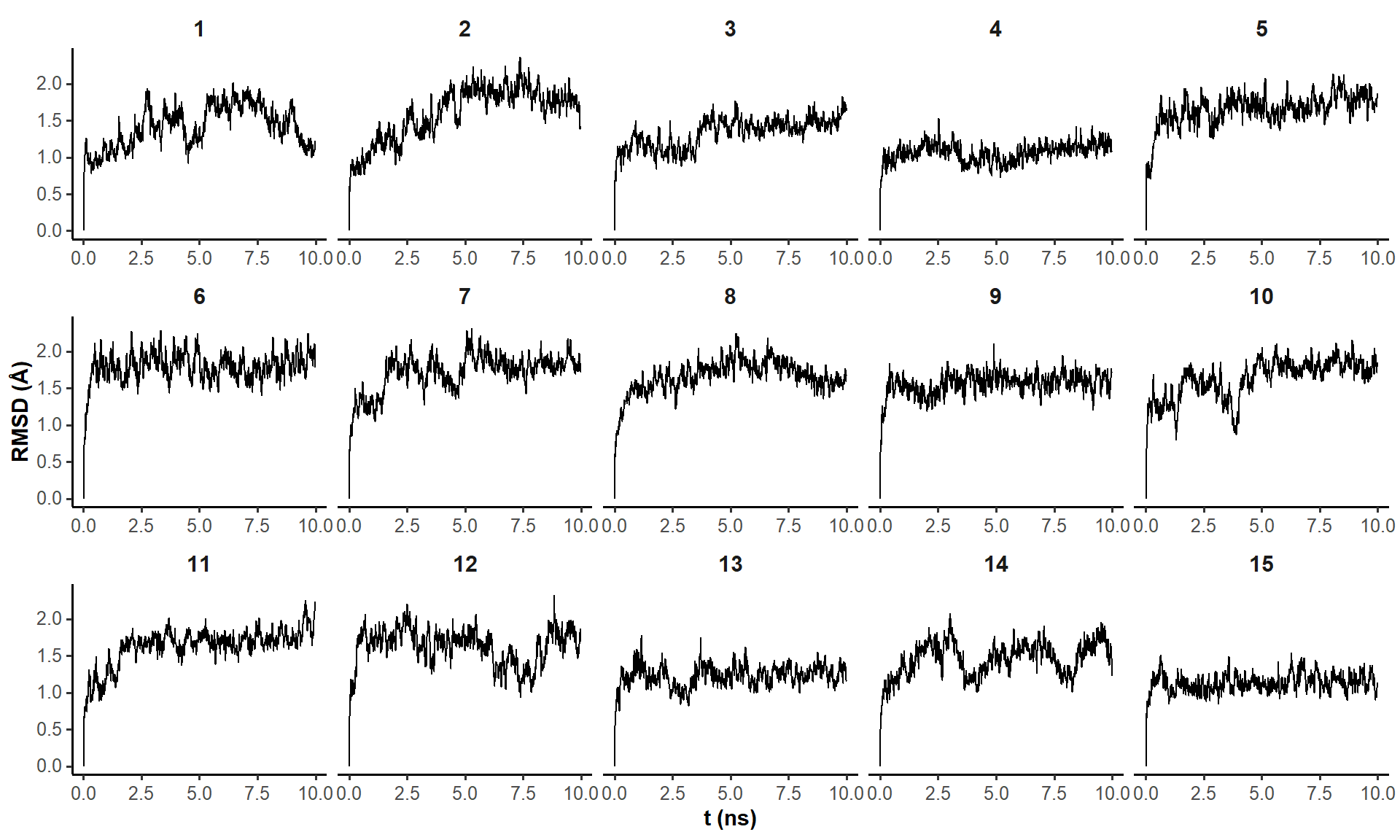
Pairwise RMSD analysis is more informative
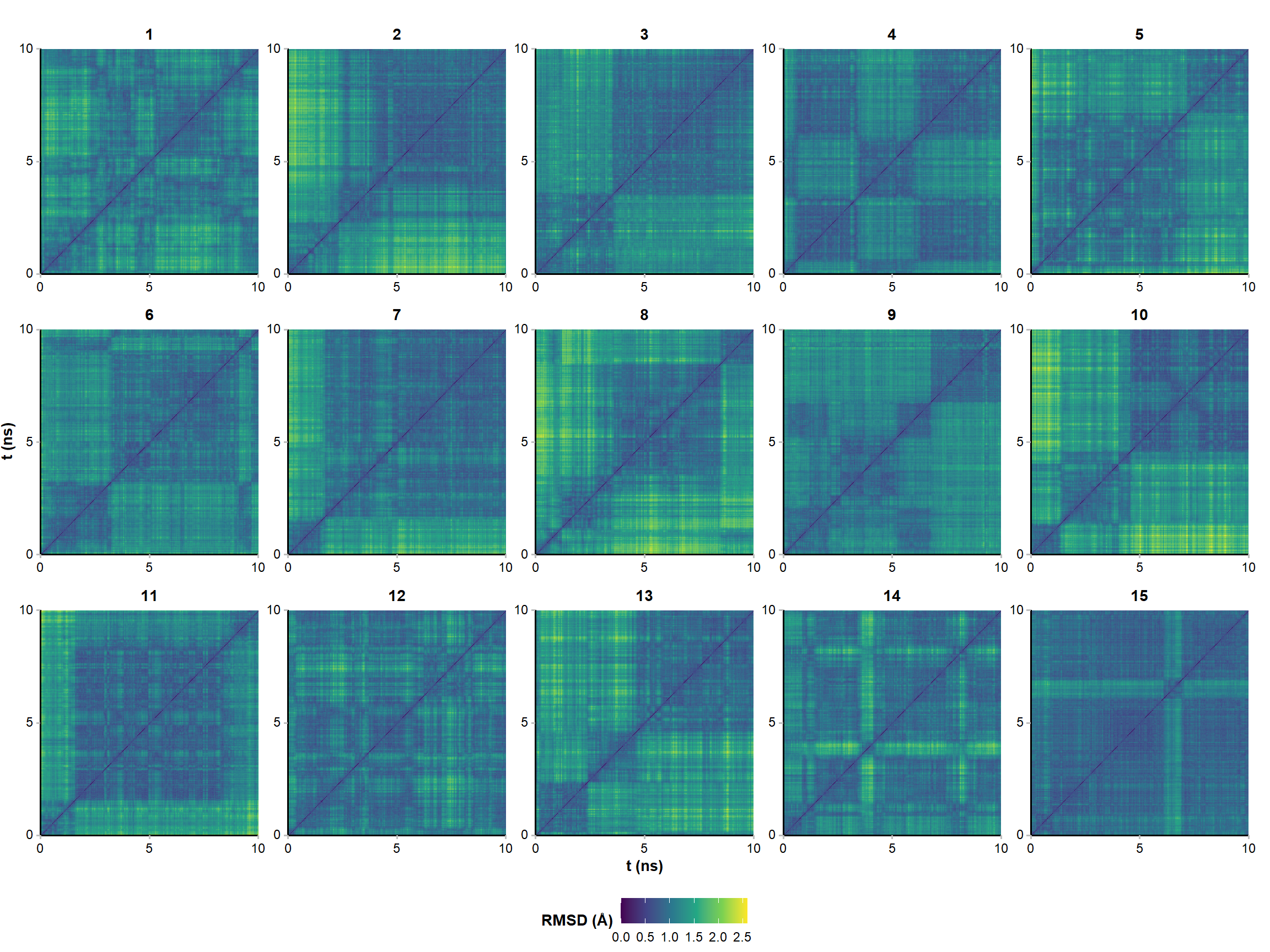
The end point of 15 runs are minimized
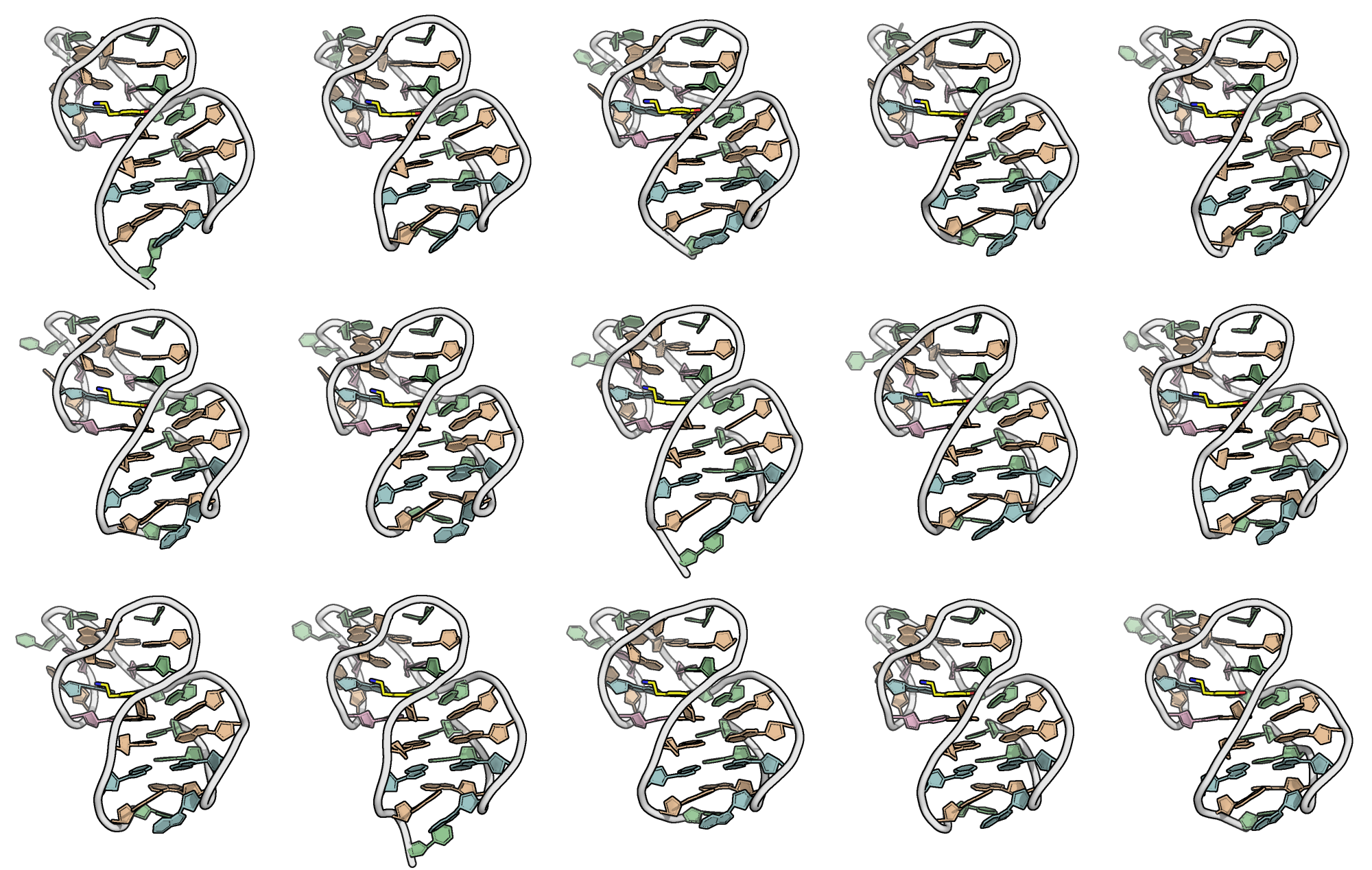
Superimposition reveals a well-defined binding pocket
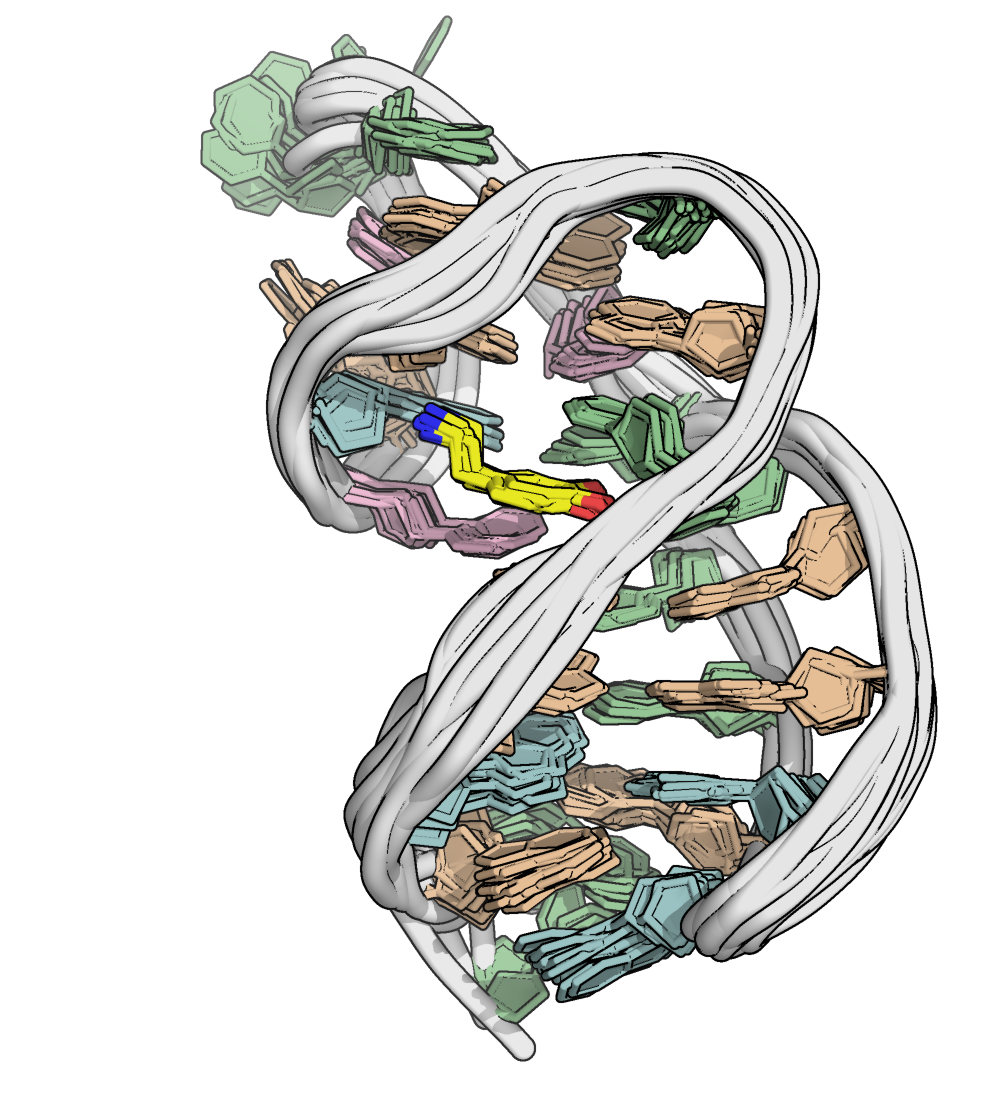
PDB: 9HIO
NMR restraints are satisfied

Exploration of the
conformational landscale
through microsecond simulations
1 µs simulation with restraints. One frame = 2 ns
Replication is necessary (at least to publish!)
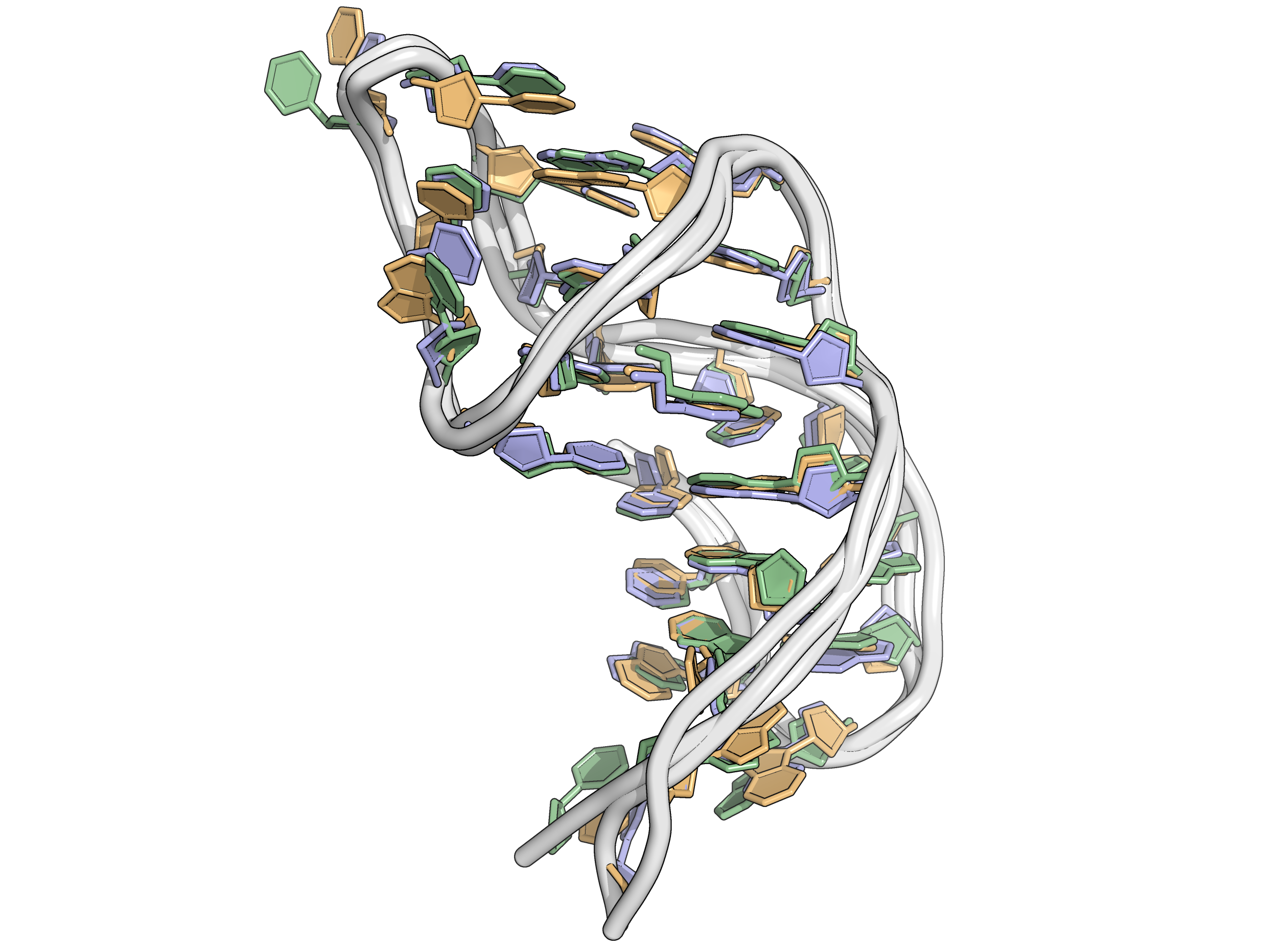
Models 3 and 9 selected as starting points for replicates of model 1 rMD
Longer simulations are stable and reproducible
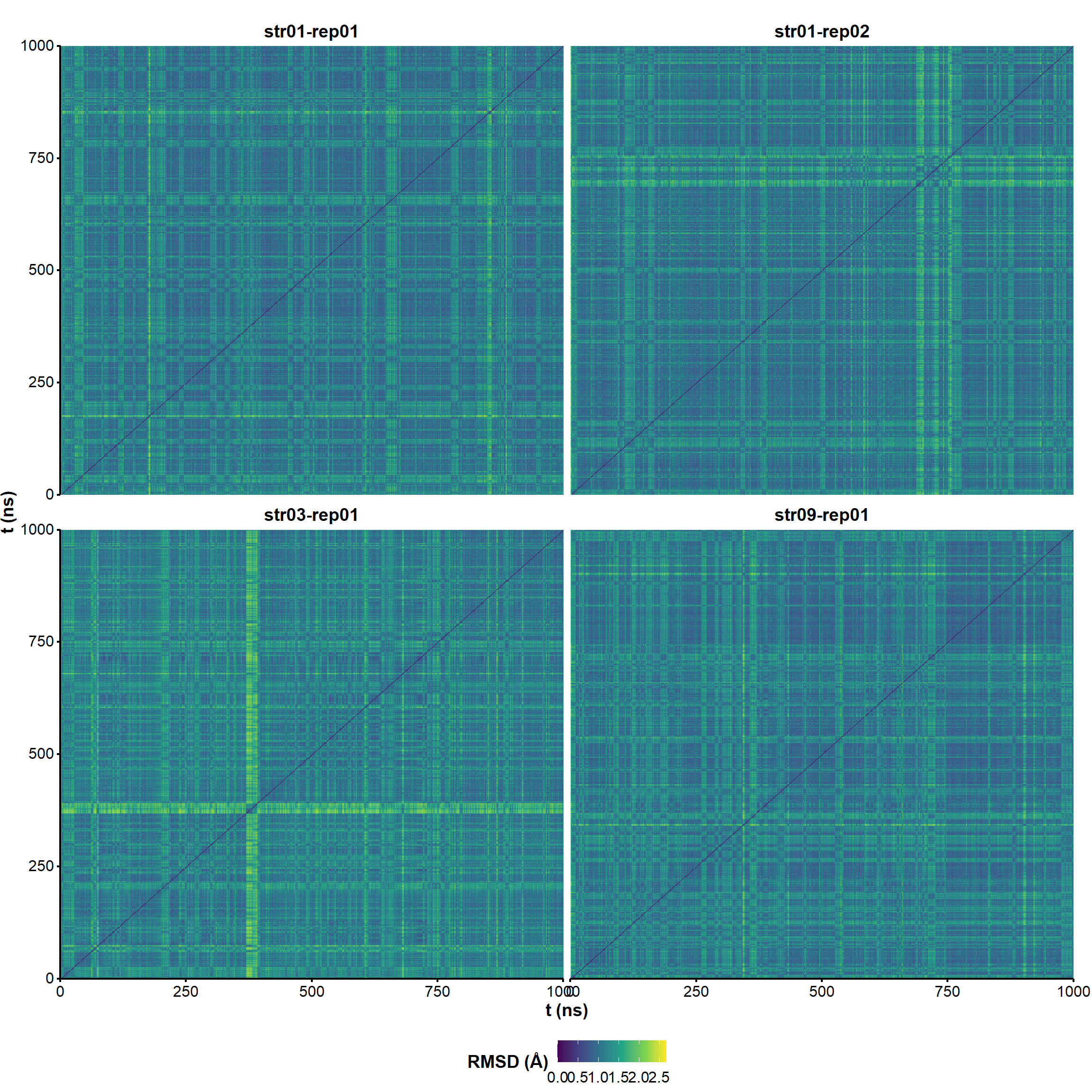
Replicates and unrestrained simulations demonstrate
the robustness of the model
Exploring non-canonical features
Non-WC base pairs and triplets
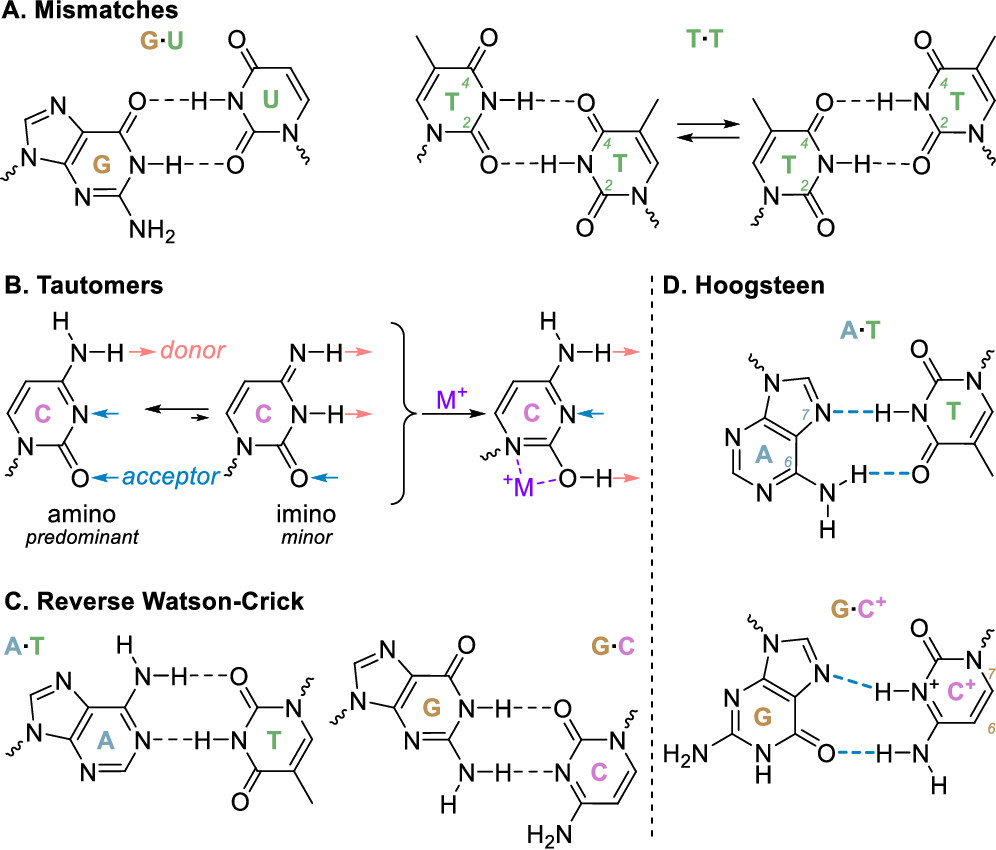
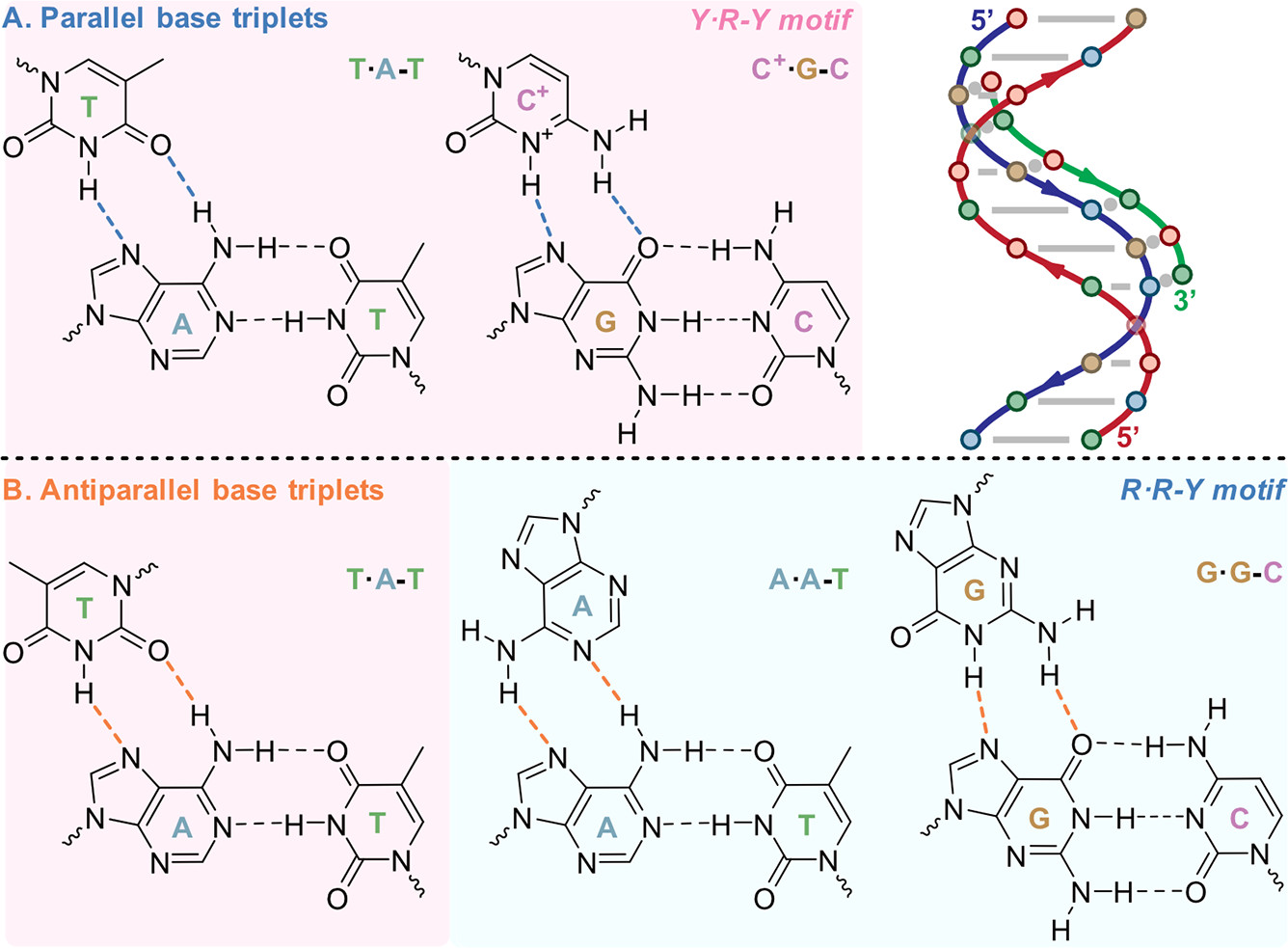
Largy et al., Chem. Rev. 2022, 122, 8, 7720–7839
Base pairing is mostly non-canonical
Automated base-pair characterization using the web API of DSSR 2.3.2 with the httr2 R package.
Only two WC base pairs are present
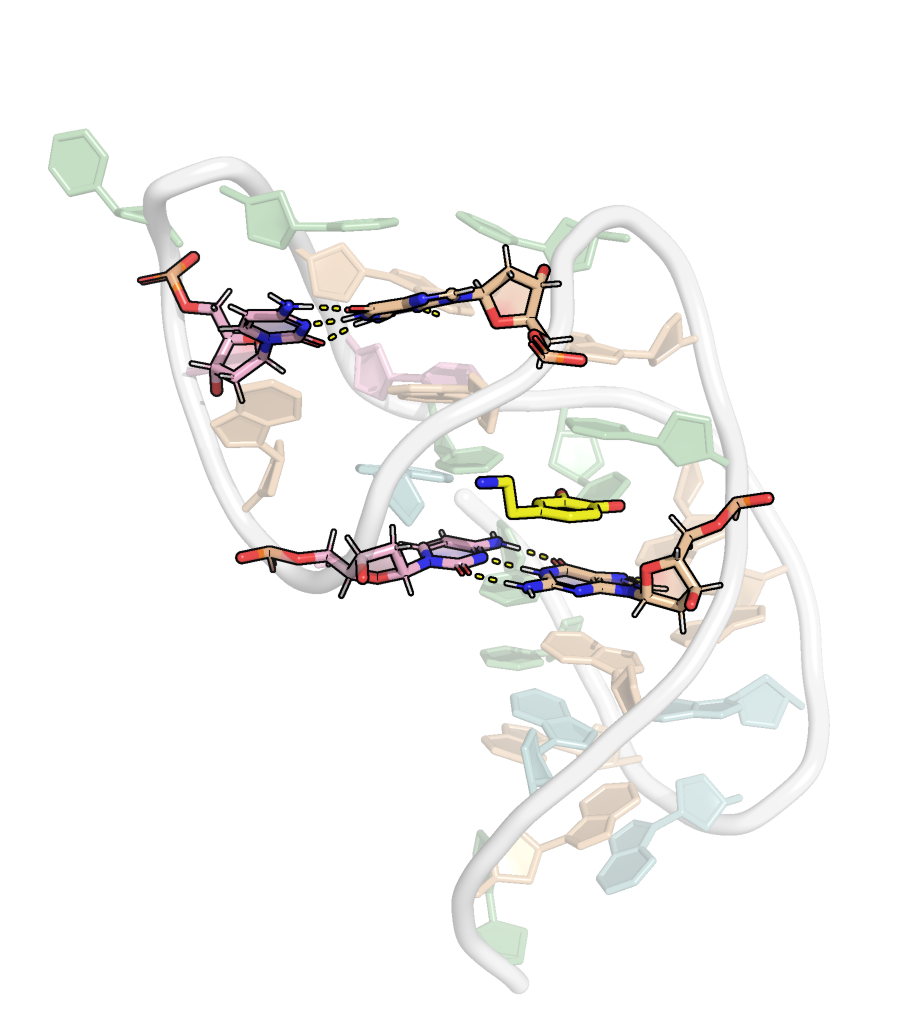
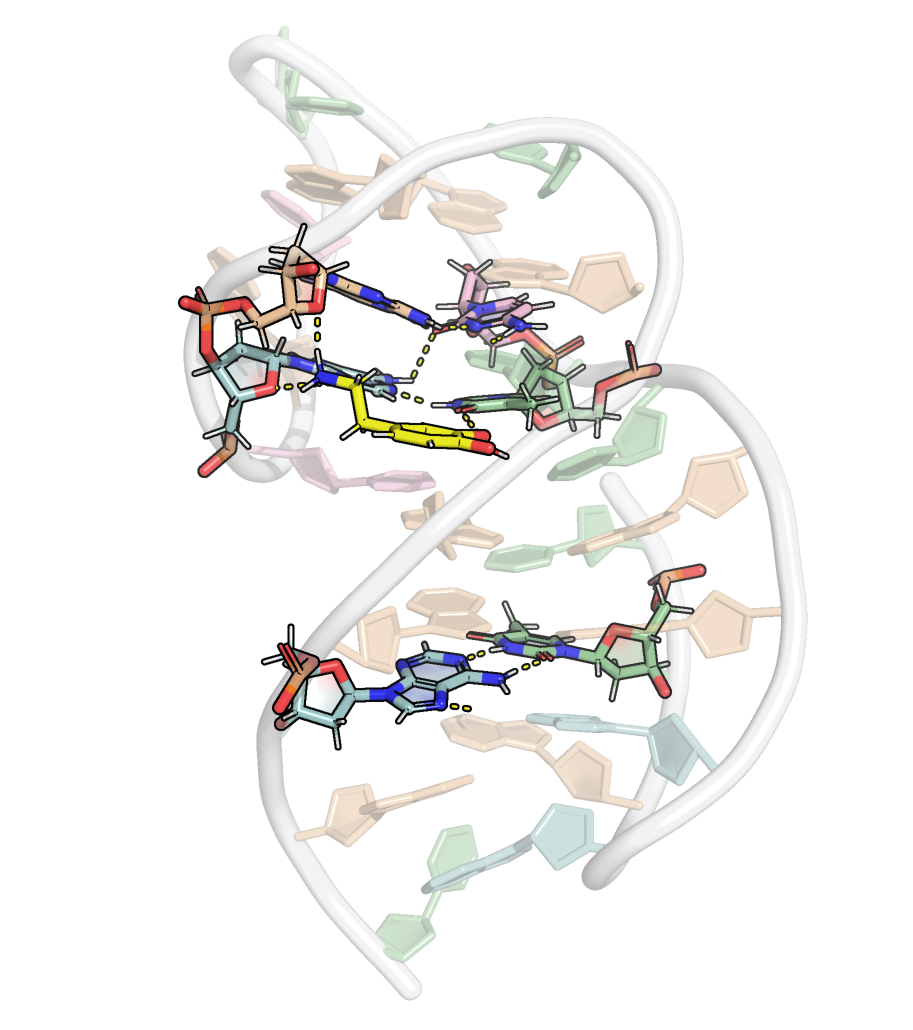
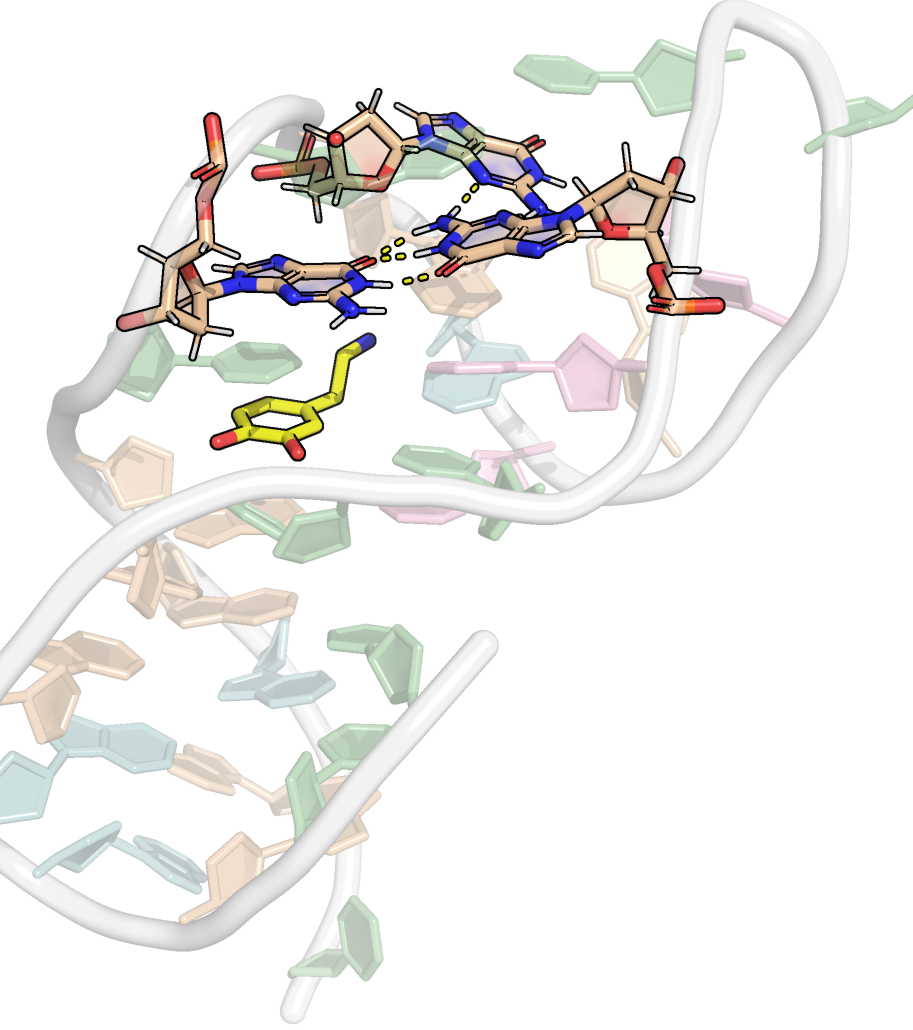
Presence of base triplets with syn glycosidic bond angle
Dihedral angles are stable along the simulation
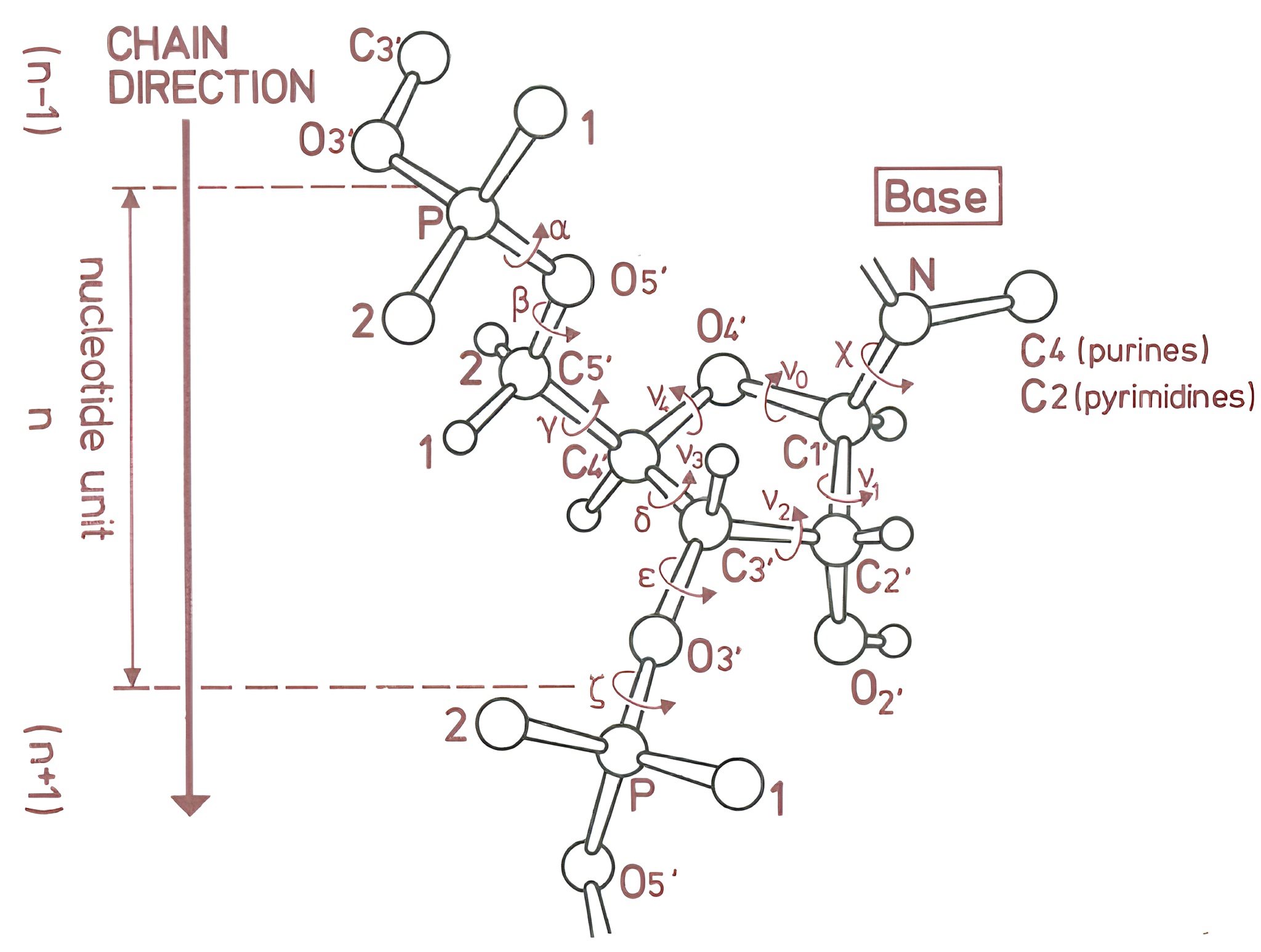
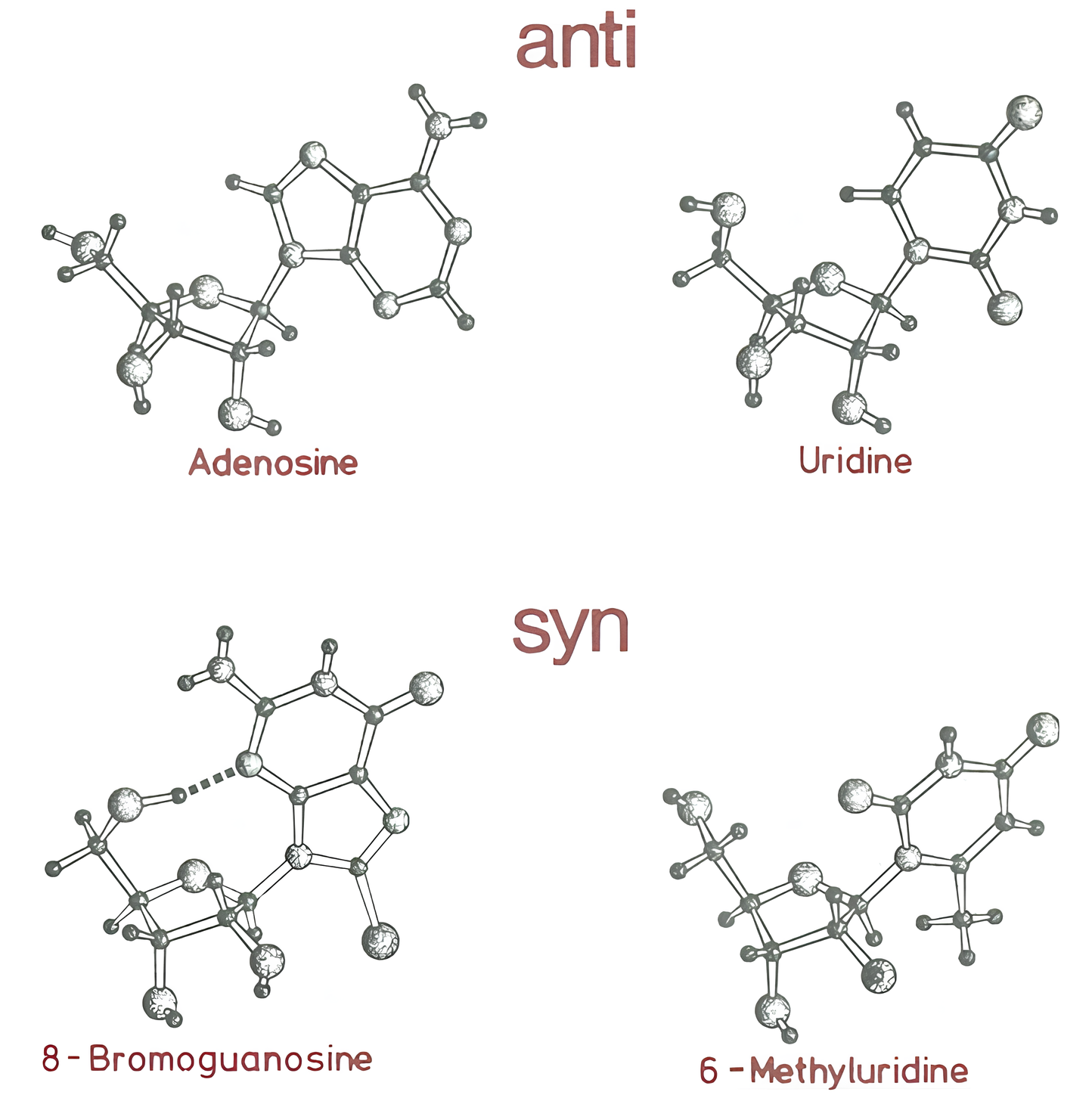


Saenger W. Principles of Nucleic Acid Structure; Springer-Verlag: New York, 1984.
Binding requires non-canonical puckers

\[\color{#F0A308}\theta_M \color{#000000}= \frac{\color{#376bd3}\nu_2}{\cos \color{#279400}P}\]
\[\tan \color{#279400}P \color{#000000}= \frac{(\color{#008594}\nu_4 \color{#000000}+ \color{#A91FB6}\nu_1\color{#000000})-(\color{#940000}\nu_3 \color{#000000}+ \color{#947E00}\nu_0\color{#000000})}{2\color{#376bd3}\nu_2\color{#000000}(\sin(\frac{\pi}{5}) + \sin(\frac{2\pi}{5}))}\]
Binding requires non-canonical puckers
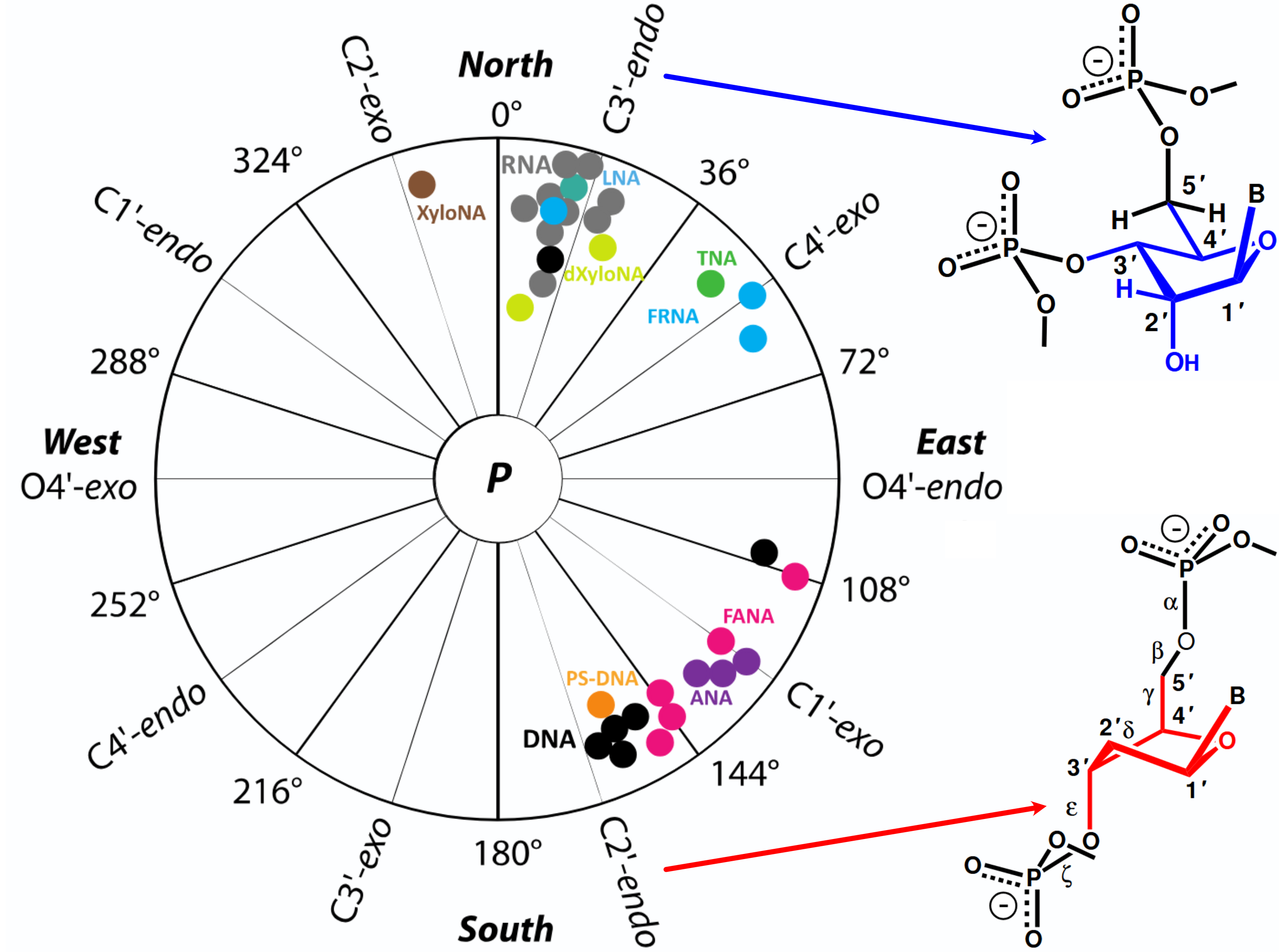
Egli, M. Sugar Pucker and Nucleic Acid Structure, 2018 11, World Scientific Publishers, 309–315. Series in Struct. Biol.
Binding requires non-canonical puckers
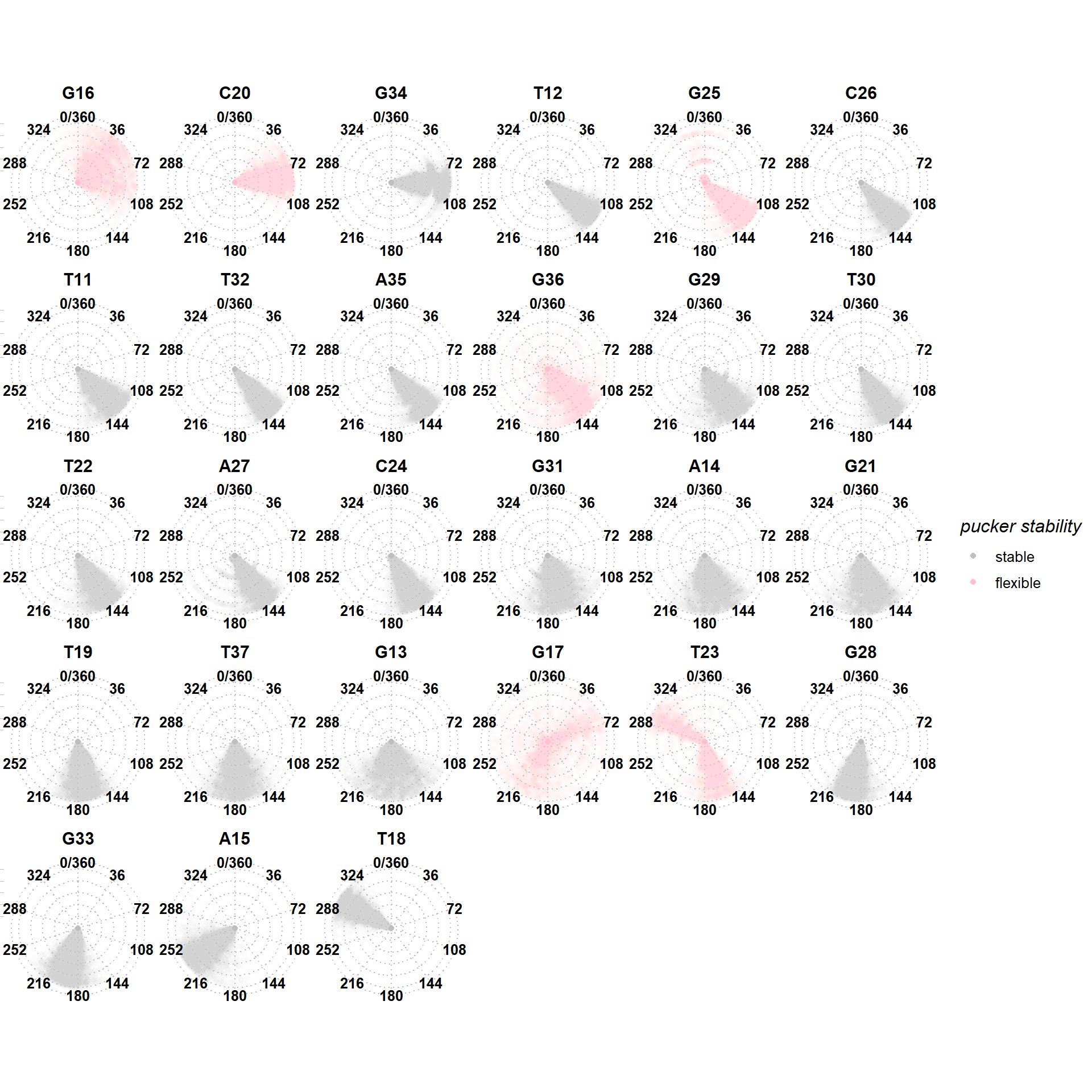
Binding requires non-canonical puckers
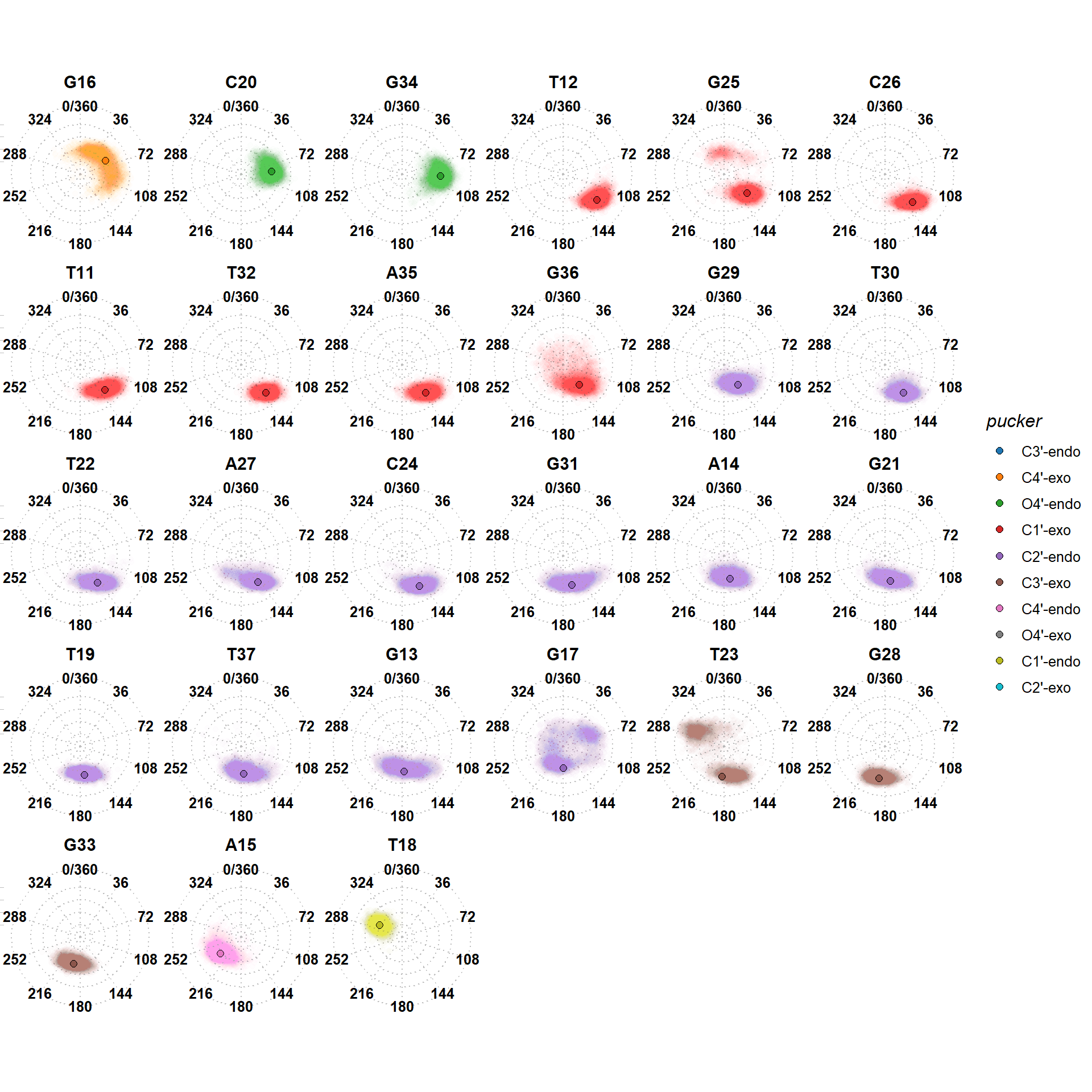

T18 adopts a C1’-endo pucker upon binding
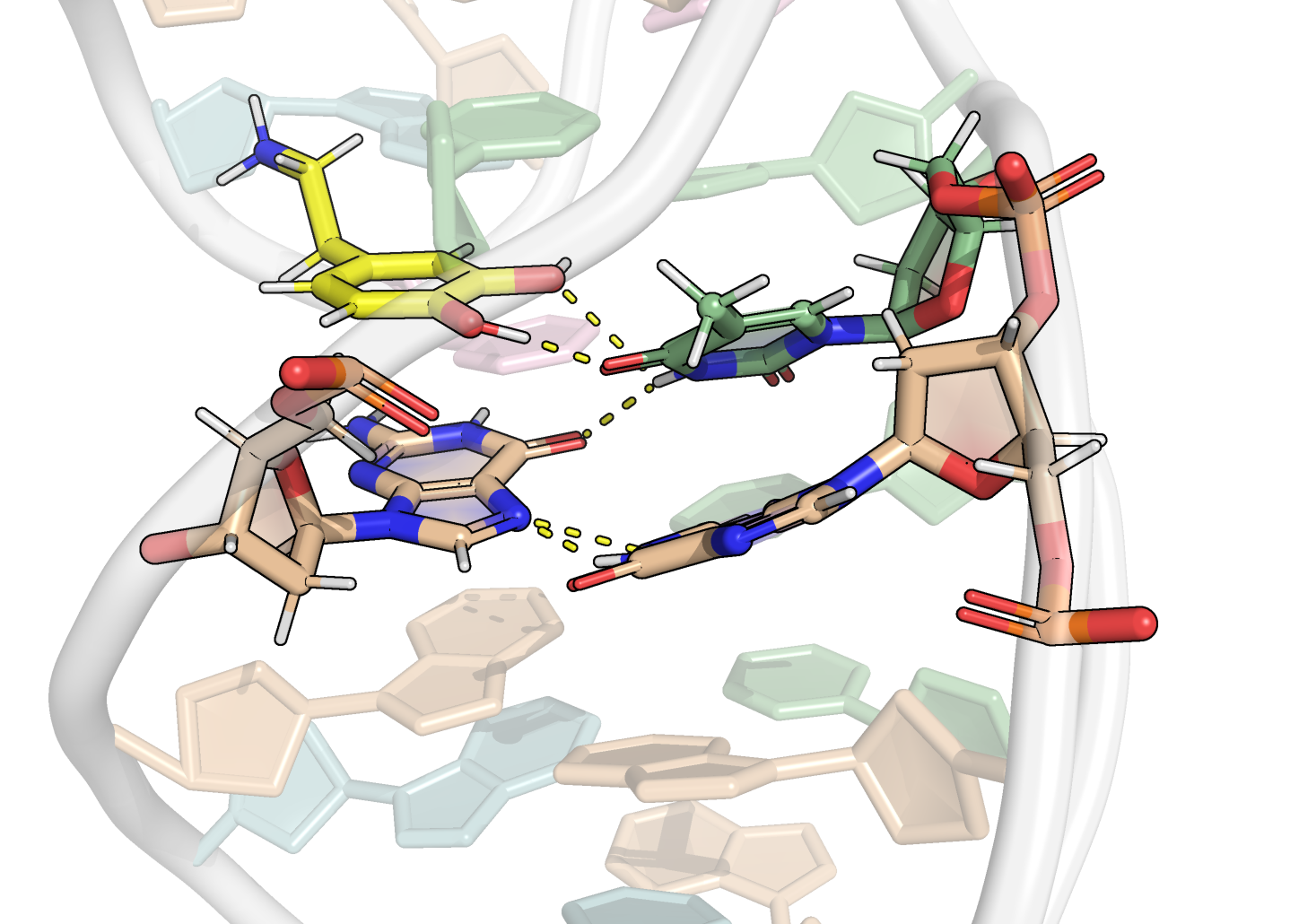
Focus on the binding site
Binding through H-bonds, electrostatic and π interactions
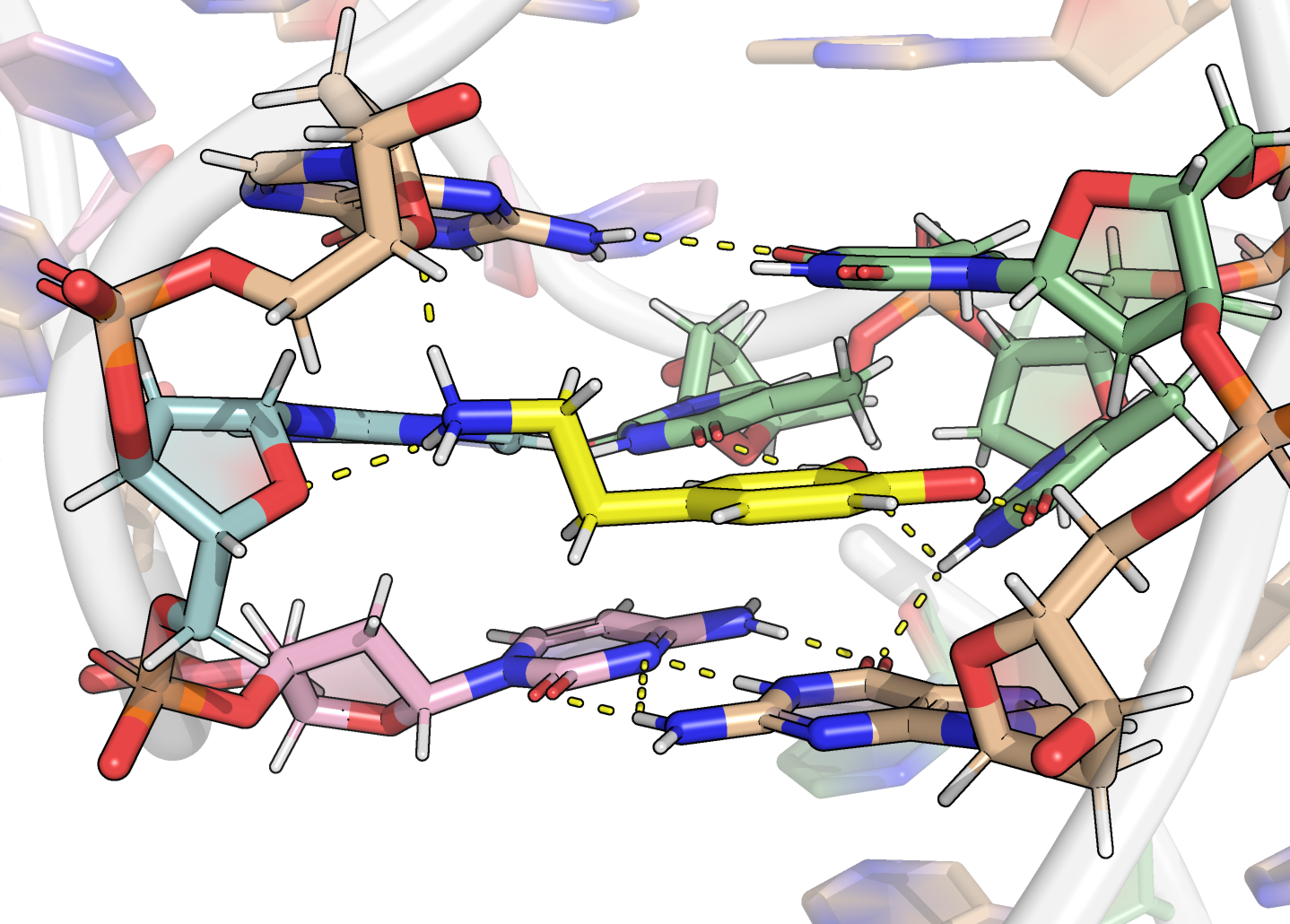
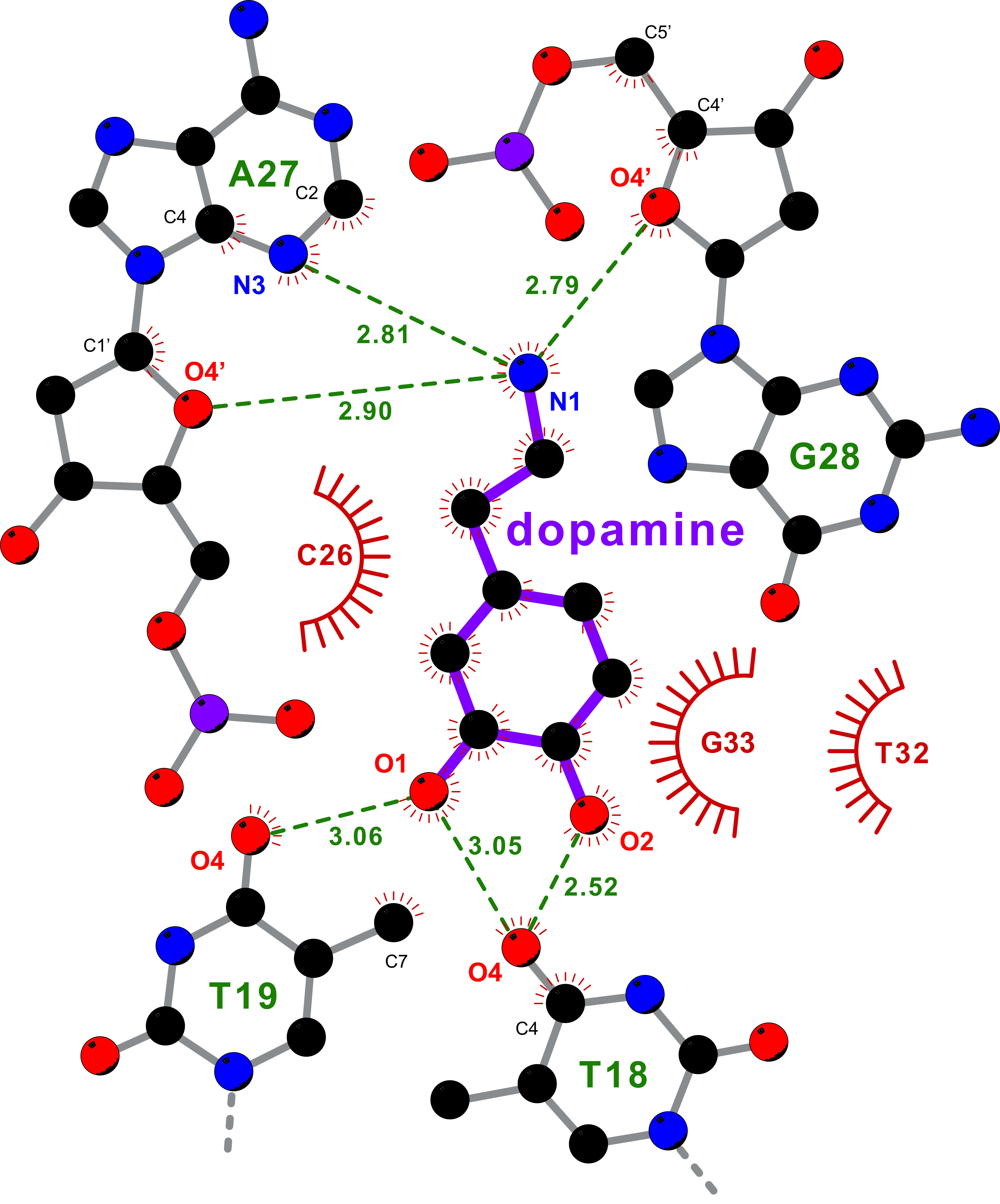
https://www.ebi.ac.uk/thornton-srv/software/LigPlus/
Restraints are necessary to refine the binding mode
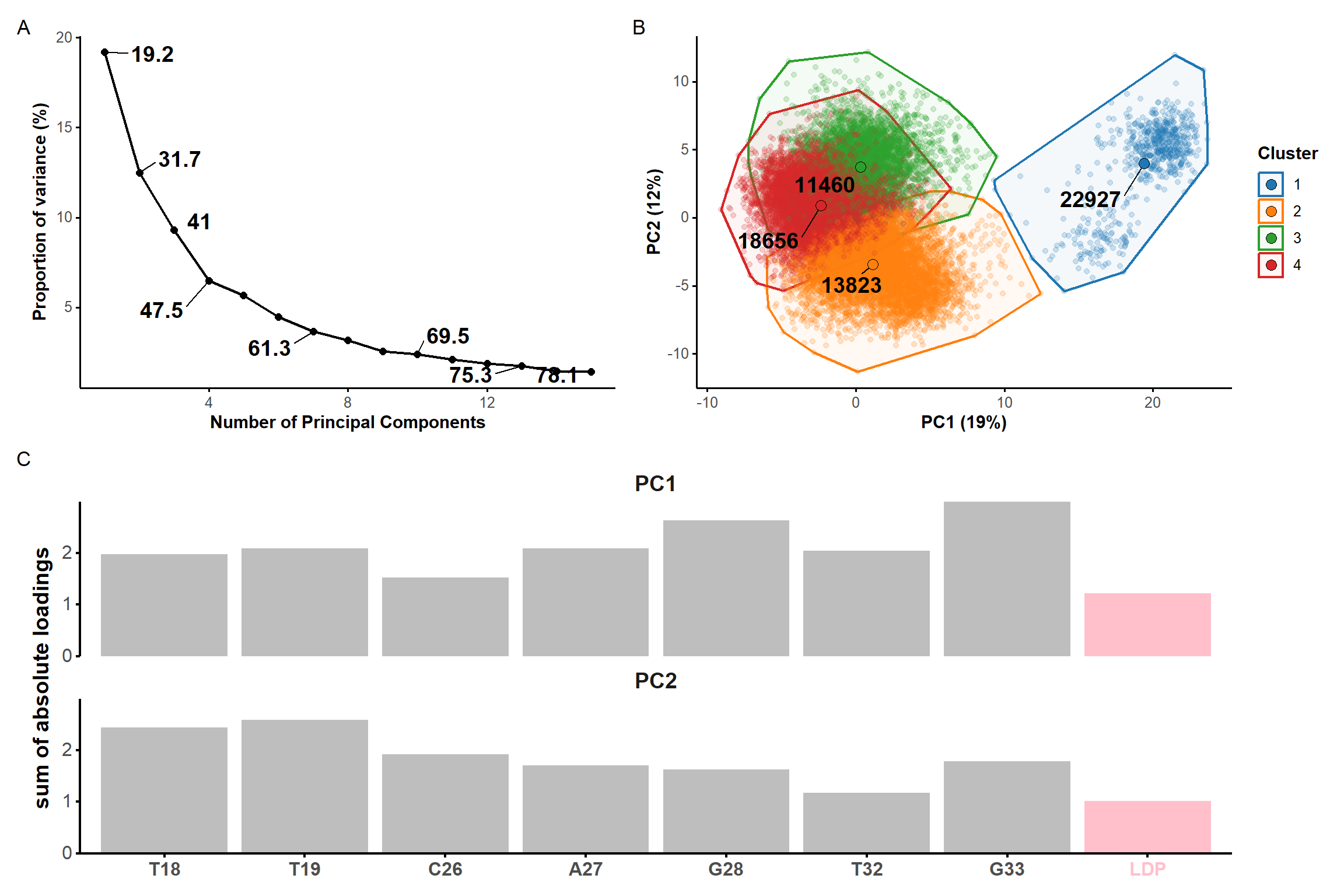
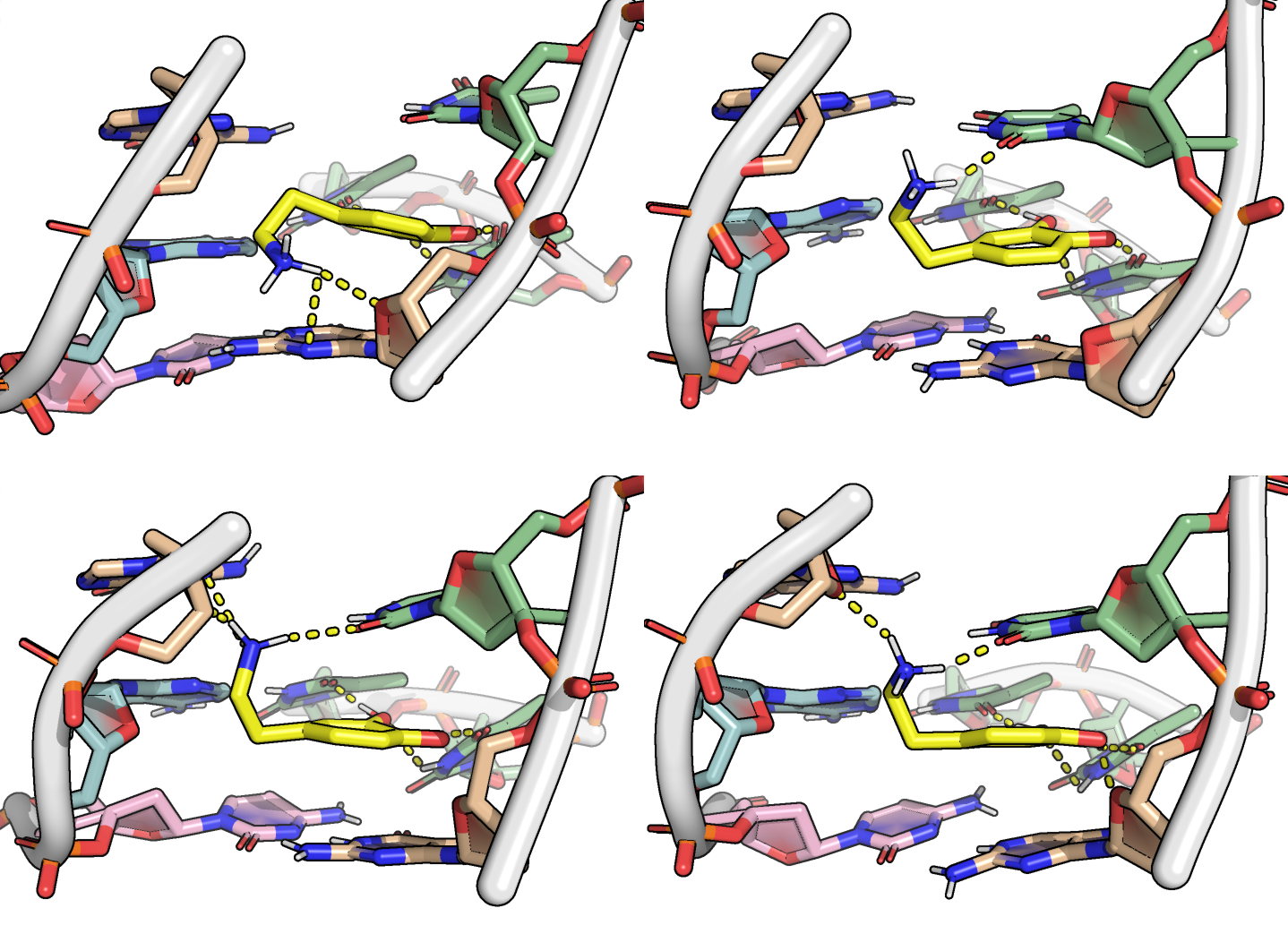
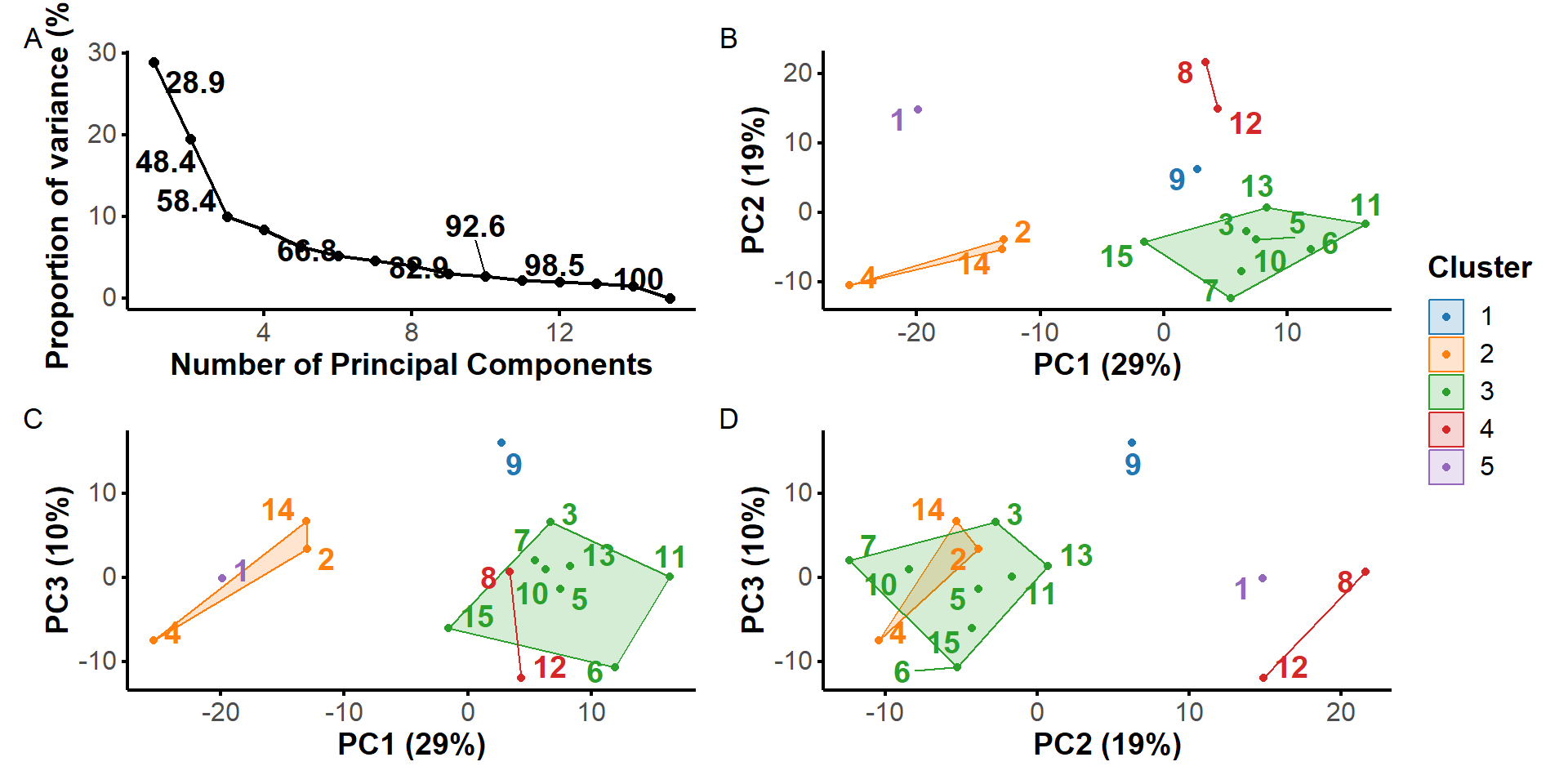
PCA performed for residues of the binding site + dopamine
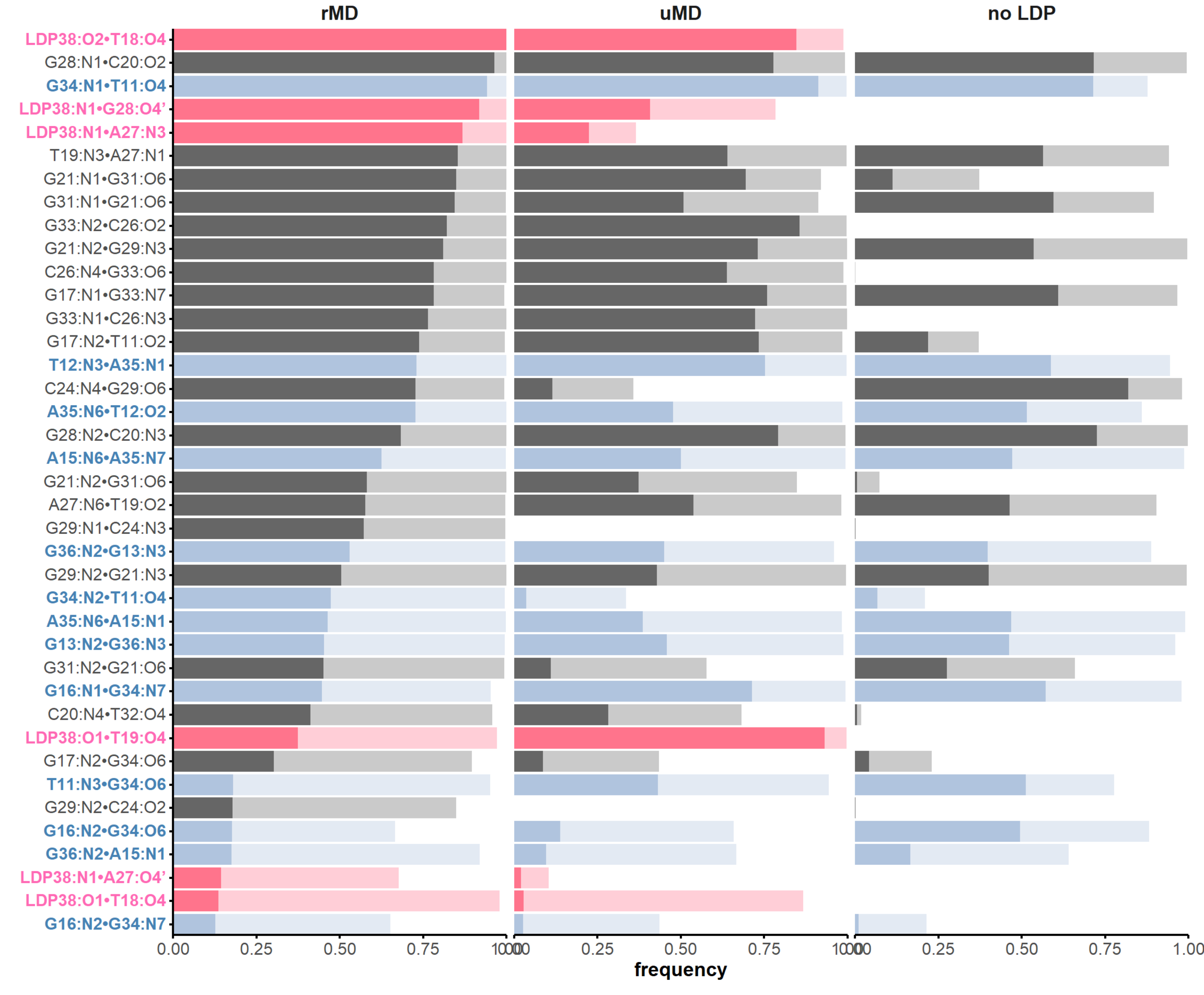
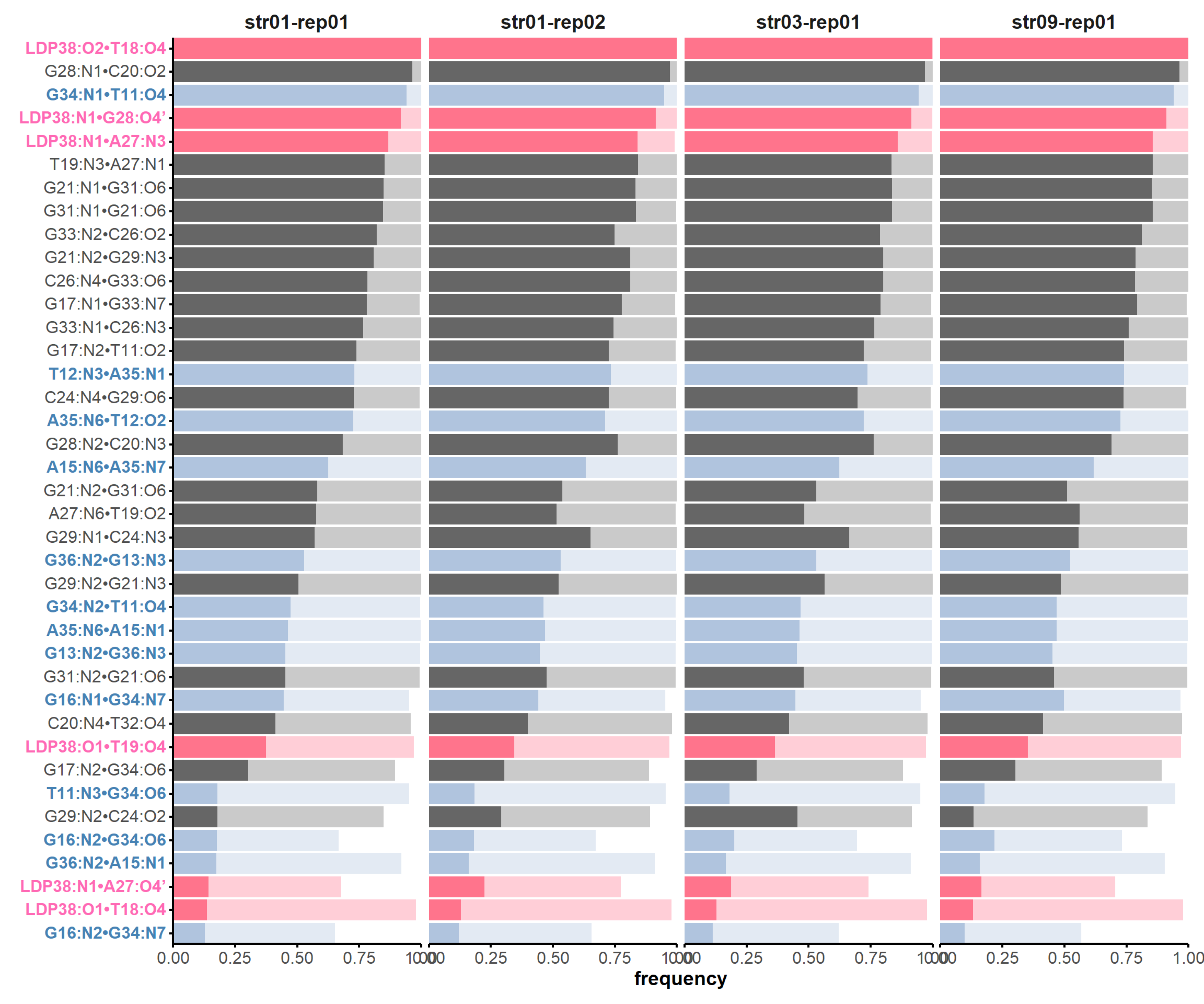
H-bonds inform structural dynamics
and model robustness
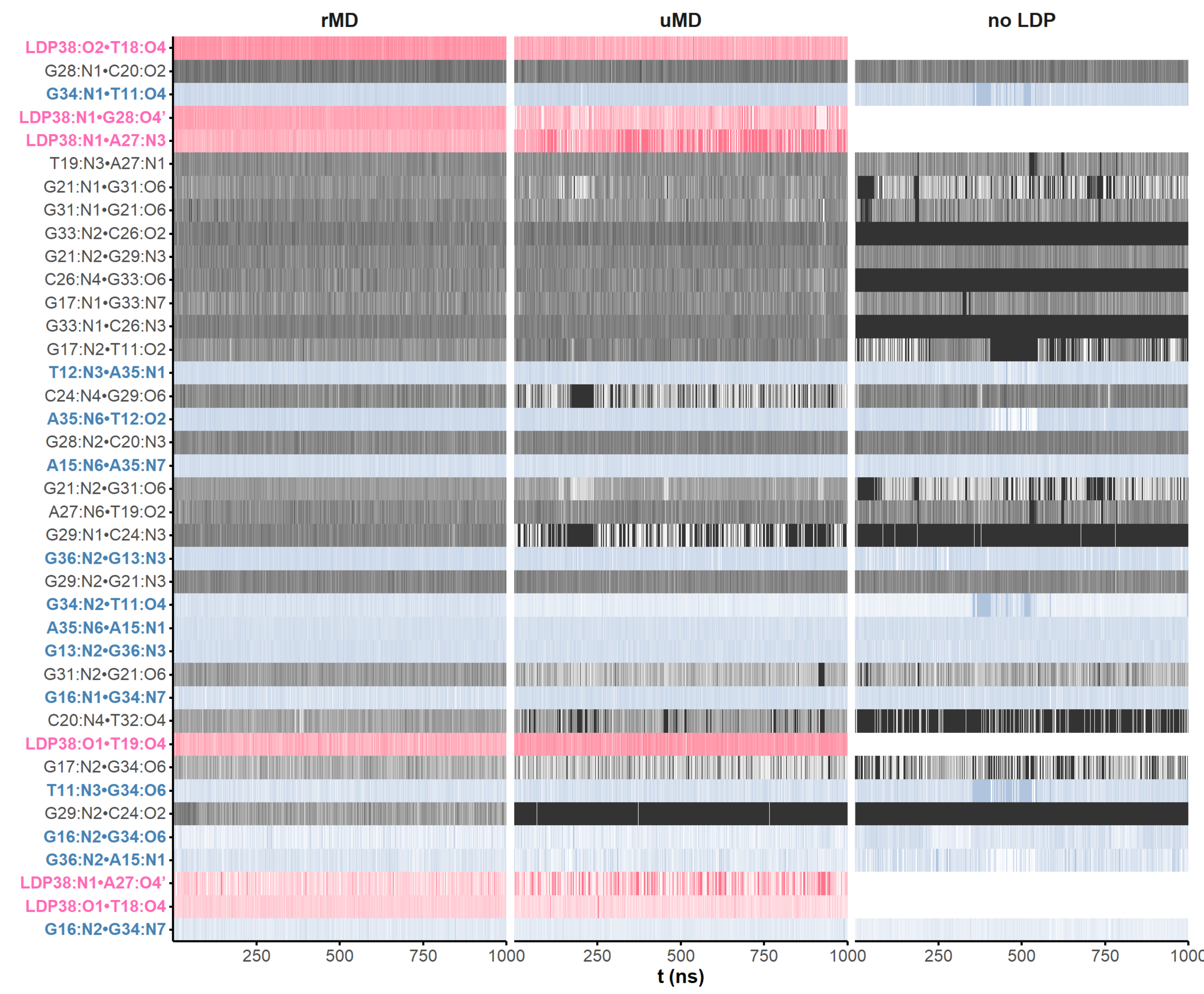
Prediction models still come short for aptamers

- 0 out of 107 submitted models with \(RMSD < 10 Å\)
- Training set is not diverse enough
- Dopamine essential for folding
Summary
- Robust MD simulations of the dopamine aptamer
- Efficient preparation workflow + replication
- GPU-enabled microsecond simulations ± NMR restraints
- Robust MD simulations of the dopamine aptamer
- Efficient preparation workflow + replication
- GPU-enabled microsecond simulations ± NMR restraints
- Semi-automated analysis of trajectories
- C++ (cpptraj), R and Python scripts,DSSR (via API) and LigPlot+
- Robust MD simulations of the dopamine aptamer
- Efficient preparation workflow + replication
- GPU-enabled microsecond simulations ± NMR restraints
- Semi-automated analysis of trajectories
- C++ (cpptraj), R and Python scripts,DSSR (via API) and LigPlot+
- Numerous diagnostics and parameters
- (pairwise) RMSD, H-bonds, dihedral angles
- Robust MD simulations of the dopamine aptamer
- Efficient preparation workflow + replication
- GPU-enabled microsecond simulations ± NMR restraints
- Semi-automated analysis of trajectories
- C++ (cpptraj), R and Python scripts,DSSR (via API) and LigPlot+
- Numerous diagnostics and parameters
- (pairwise) RMSD, H-bonds, dihedral angles
- Very non-canonical elements are necessary for folding/binding
- Robust MD simulations of the dopamine aptamer
- Efficient preparation workflow + replication
- GPU-enabled microsecond simulations ± NMR restraints
- Semi-automated analysis of trajectories
- C++ (cpptraj), R and Python scripts,DSSR (via API) and LigPlot+
- Numerous diagnostics and parameters
- (pairwise) RMSD, H-bonds, dihedral angles
- Very non-canonical elements are necessary for folding/binding
- Prediction model have a long way to go for aptamers
Experimental data is necessary to solve aptamer structures
- Robust MD simulations of the dopamine aptamer
- Efficient preparation workflow + replication
- GPU-enabled microsecond simulations ± NMR restraints
- Semi-automated analysis of trajectories
- C++ (cpptraj), R and Python scripts,DSSR (via API) and LigPlot+
- Numerous diagnostics and parameters
- (pairwise) RMSD, H-bonds, dihedral angles
- Very non-canonical elements are necessary for folding/binding
- Prediction model have a long way to go for aptamers
Acknowledgments
- Canadian resources 🍁
- Cameron Mackereth | Prism Team, ARNA, INSERM U1212, CNRS UMR 5320, U. Bordeaux
- Philip E. Johnson | Department of Chemistry, York University, Toronto, ON, Canada
- Computational resources 💻
- DOREMI CALI v3 cluster of the Mésocentre de Calcul Intensif Aquitain (U. Bordeaux)